최소 단어 이상 선택하여야 합니다.
최대 10 단어까지만 선택 가능합니다.
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
NTIS 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
DataON 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Edison 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Kafe 바로가기
| 주관연구기관 | 고려대학교 산학협력단 |
|---|---|
| 연구책임자 | 전태훈 |
| 참여연구자 | 홍기창 , 이영식 , 한지혜 , 배준범 , 최창용 , 최상필 , 이승재 , 그외 다수 |
| 보고서유형 | 최종보고서 |
| 발행국가 | 대한민국 |
| 언어 | 한국어 |
| 발행년월 | 2015-03 |
| 과제시작연도 | 2011 |
| 주관부처 | 농촌진흥청 |
| 연구관리전문기관 | 농촌진흥청 Rural Development Administration |
| 등록번호 | TRKO201500010192 |
| 과제고유번호 | 1395022181 |
| DB 구축일자 | 2015-07-11 |
Ⅳ. 연구개발결과
PMWS 발병 방지를 위한 면역조절 기전 구명 및 이를 이용한 질병저항성 활용 기술 개발을 위해 PCV2 ORF와 상호작용하는 면역 숙주세포 내 표적 분자를 동정하고 하부신호를 탐색하여 발병메커니즘을 확인하였다. 또한, PCV2 ORF 발현에 의해 변화되는 숙주세포의 마이크로 RNA와 그 표적 유전자를 동정하여 발현조절에 따른 PCV2 감염저항성조절 기반기술을 개발하였다. PCV2 증식관련 측정형질을 발굴하여 PCV2증식억제관련 후보유전자를 선정하였고, 후보유전자 내 신규 염기다형성을 발굴하였다. 또한, 연관
Ⅳ. 연구개발결과
PMWS 발병 방지를 위한 면역조절 기전 구명 및 이를 이용한 질병저항성 활용 기술 개발을 위해 PCV2 ORF와 상호작용하는 면역 숙주세포 내 표적 분자를 동정하고 하부신호를 탐색하여 발병메커니즘을 확인하였다. 또한, PCV2 ORF 발현에 의해 변화되는 숙주세포의 마이크로 RNA와 그 표적 유전자를 동정하여 발현조절에 따른 PCV2 감염저항성조절 기반기술을 개발하였다. PCV2 증식관련 측정형질을 발굴하여 PCV2증식억제관련 후보유전자를 선정하였고, 후보유전자 내 신규 염기다형성을 발굴하였다. 또한, 연관성분석을 통해 PCV2증식수준 최적 유전자형을 선정하였고, 돼지 SNP chip 분석을 통해 PCV2증식관련 유용 SNP도 발굴하였다. 뿐만 아니라, 자돈 생존폐사와 관련된 SNP를 대량 발굴하여 200개의 기능성 후보유전자를 검색하고, 자돈소모성 질환과 관련된 원인유전자 및 기능을 규명하는데 정보를 제공하였다. 돼지 MHC(SLA) 타이핑을 실시하여 면역관련 유전자의 유전자형 분석을 실시하였다. 이에 따라 viral epitope 결합력 분석 시스템을 개발하여 질병저항성 품종개발에 필요한 SLA class I 및 II의 대립 유전자 데이터베이스를 확장하였다.
< Development of genetic markers related to the resistance to PMWS by dissecting the molecular mechanism of PCV2 and host cell interactions >
Porcinecircovirus type 2 (PCV2) is the primary causative agent of postweaning multisystemic wasting syndrome (PMWS), which leads to serious economic losses
< Development of genetic markers related to the resistance to PMWS by dissecting the molecular mechanism of PCV2 and host cell interactions >
Porcinecircovirus type 2 (PCV2) is the primary causative agent of postweaning multisystemic wasting syndrome (PMWS), which leads to serious economic losses in the pig industry worldwide. The mechanism of pathogenicity associated with PCV2 infection is still not fully understood. Nevertheless, the fact that large amounts of proinflammatory cytokines within lymphoid tissues are released during the early stage of PCV2 infection may induce chronic inflammatory responses followed by the destruction of lymphoid tissues. However, how PCV2 infection causes an excessive inflammatory response in the host immune system during the early stage of PCV2 infection is still not elucidated. In this study, we show that direct interaction between the PCV2 ORF3 and regulator of G protein signaling 16 (RGS16) within the cytoplasm of host cells leads to ubiquitin-mediated proteasomal degradation of RGS16. Facilitated degradation of the RGS16 by PCV2 ORF3 further enhances NFkB translocation into the nucleus through the ERK 1/2 signaling pathway and increased IL-6 and IL-8 mRNA transcripts. Consequently, more severe inflammatory responses and leukocyte infiltration occur around host cells. Also, we analyzed the expression of C1qBP that other target protein of PCV2 ORF. The results show that the expression of C1qBP is increased in PCV2 ORF2 overexpressed host cells and phagocytosis activity is also increased. And, we employed Solexa deep sequencing technology to determine which cellular miRNAs were differentially regulated after expression of each of three PCV2-encoded ORFs in host cells. We identified 51 ORF1-regulated miRNAs, 74 ORF2-regulated miRNAs, and 32 ORF3-regulated miRNAs that differed in abundance compared to the control. Among the potential target genes of ORF-regulated miRNAs, two genes encoding proteins that are known to interact with PCV2-encoded proteins, zinc finger protein 265 (ZNF265) and RGS16, were selected for further analysis. We provide evidence that ZNF265 and RGS16 are direct targets of miR-139-5p and let-7e, respectively, which are both down-regulated by ORF2.
< Identification of PCV2 proliferation resistance DNA marker in pig >
The goal of this project is identification of PCV2 proliferation resistance DNA marker in pig. Therefore, we decided a major trait of PCV2 proliferation measurement on a PCV2 genomic DNA quantity in porcine peripheral blood. PCV2 genomic DNA quantity (Viral load) in porcine peripheral blood from 200 pigs were quantified using qPCR. We decided two candidate genes, Porcine Regulator of G protein signaling 16 (RGS16) and MYC, which are interact with PCV2 through the information about PCV2 related genes. 22 single nucleotide polymorphisms (SNPs) were founded in RGS16 and 1 SNP and 1 InDel were founded in MYC gene using pooling DNA sequencing. However, the SNP and InDel in MYC were identified in Berkshire breed. So, we could not type the genotype of SNP in MYC because, we had Yorkshire breed samples. Genotypes of each SNPs from 142 Yorkshire pigs were typed using PCR-RFLP and direct sequencing methods. Three SNPs (SNP2, 8, 11) showed significant correlation between genotypes and PCV2 viral load (each P = 0.022, P = 0.033, P = 0.027). Additionally, we discovered another candidate gene, DNAJC6, using Genome wide association study (GWAS) and identified a new SNP and genotyped about 142 Yorkshire pigs using PCR-RFLP. The association study between genotypes in DNAJC6 SNP and PCV2 viral load was shown significant correlation (P < 0.040). For effectiveness verification of DNA markers, we quantified PCV2 viral load, meat quality traits, and lean meat production traits from 417 Yorkshire pigs in normal farms. Association study showed significant correlation between the genotype of SNP2 in RGS16 and lymphocyte count (P < 0.042). Also, the SNP2 in RGS16 showed significant correlation with Lightness (L*) (P < 0.013). The SNP11 in RGS16 significantly correlated with backfat thickness (P < 0.026), pH45min (P < 0.031), and Lightness traits (L*) (P < 0.002).
< Detection of SNPs related to blood immune components and survival/death of piglets >
The importance of the genes related to SLA, immune response and resistance to PMWS disease in pigs is increasing due to the big loss of swine production caused by infectuous disease. A set of 40 significant SNPs were detected by genome-wide association analysis using high throughput porcine SNP chips in an F2 cross population between Korean native pigs and Yorkshire. About 40 functional candidate genes were found which were located near the significant SNPs for the blood immune components. A set of 449 SNPs were detected for survival & death of piglets using the Illumina porcine 64K SNP chip using the samples of 72 live and dead piglets, and about 200 functional candidate genes were found near the significant SNPs. The SNPs would be provided as SNP resources to find causal genes for resistance to PMWS.
< Characterization and excavating of the immune response related genes for improving the tolerance on wasting disease in the piglet >
Unlike the long-term information has been accumulated human and mouse, pig Major histocompatibility complex (MHC) and the study of the association between disease-resistant case can not be made a lot of research because of the technical limitations debilitating disease and swine MHC allele-specific reactivity differences associated the need for research to identify significant. To this end, the case of swine SLA information stored on the allele is very necessary and it is possible to develop epitope prediction program for swine MHC and TCR a future based on it. Research on swine MHC (SLA) is a restriction fragment length polymorphism (RFLP) haplotype constructed by using the method compared with the HLA (. Chardon et al, 1985) class I and class II haplotype analysis for the occasion and these genes a recombinant haplotype for the analysis (Chardon et al., 1985; Lie et al., 1987; Singer et al., 1987; Sachs et al., 1988; Martens et al., 2003) are primarily been used came. DNA molecular marker-based selection through the pig genome using DNA markers by combining genetic techniques (MAS, marker assisted selection) in situations that are less precise molecular transition is known about the cause of the excessive growth inhibition and immune-deficient PCV2 far and to develop and provide a fundamental solution of PMWS. Piglets through an analysis using the half of the money intended for commercial analysis and the main body of the SLA genes immunomodulatory genes are expected to be the most important gene in relation to the development of useful DNA markers with wasting disease resistance, and resistance to common diseases associated with existing reported to the usefulness gene (FUT1, Mx1, Nramp1) and if there is a relationship between rates and livestock wasting disease requires a further study of the patent avoidance technologies. For the analysis of SLA genotype and haplotype, two groups of 300 early died and 300 normal piglets were typed using genomic DNA sequence based typing (GSBT; developed by our previous research). Based on the GSBT results, the haplotype of both groups were analyzed through the pedigree analysis. To analyze the genotype of non-SLA immunomodulatory genes, between early died group and normal group, piglets diarrhea E. coli associated resistance and the virus-related genes such as FUT1, TNFA, TNFB, Mx1, Nramp1, TLR, MUC4, SLC12A8, TFRC, MYD88, pBD, play an important role in the regulation of immune response and defense of foreign antigens. For the excavation of SNPs, SNPs was confirmed by DNA sequencing after amplification by PCR using gene specific primer sets. The minor allele frequencies of SNP alleles among the excavated using a PCR-RFLP, direct sequencing or Snapshot way to target the SNP should only be carried out at least 10% of SNP typing. Analysis of statistical significance for the SLA haplotype between early died and normal piglets performed using POPGENE 1.31 software for the observed and expected allele frequency measurement. Statistical analysis to estimate the SNP genotype and allele effects on piglet survival significance tests to be conducted by chi-square test (chi-square test). For multiple test correction, a Bonferroni correction applied. Newly excavated SLA alleles were added to the largest international SLA database. Developed database characterized on a specific pathogen pig MHC epitope (epitope) and related vaccines against swine MHC alleles with disease allele. SLA alleles can be used as a useful data for the development of information-based disease-resistant varieties.
 표
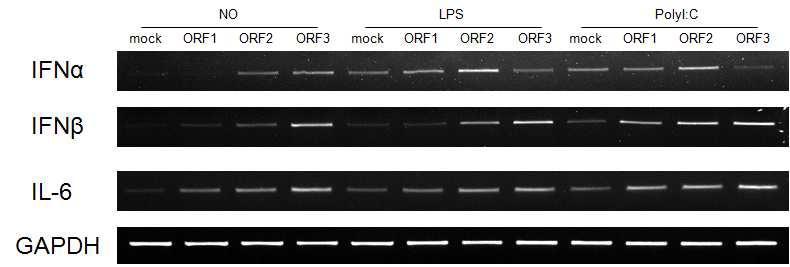
숙주세포(PK15 세포) 내 PCV2의 ORF1, ORF2, ORF3 발현과 IFN-alpha, IFN-beta, IL-6의 발현량과의 상관 관계. 감염된 PK15 세포에 toll-like receptor (TLR)의 주요 ligand인 LPS (TLR2)와 poly(I:C) (TLR3)를 처리한 후, 변화하는 IFN-alpha, IFN-beta, IL-6의 발현량을 RT-PCR로 분석하였음. No, TLR ligand를 처리하지 않은 시료; LPS, LPS를 처리한 시료; Poly(I:C), poly(I:C)를 처리한 시료; mock, empty vector only.
표
숙주세포(PK15 세포) 내 PCV2의 ORF1, ORF2, ORF3 발현과 IFN-alpha, IFN-beta, IL-6의 발현량과의 상관 관계. 감염된 PK15 세포에 toll-like receptor (TLR)의 주요 ligand인 LPS (TLR2)와 poly(I:C) (TLR3)를 처리한 후, 변화하는 IFN-alpha, IFN-beta, IL-6의 발현량을 RT-PCR로 분석하였음. No, TLR ligand를 처리하지 않은 시료; LPS, LPS를 처리한 시료; Poly(I:C), poly(I:C)를 처리한 시료; mock, empty vector only.
 표
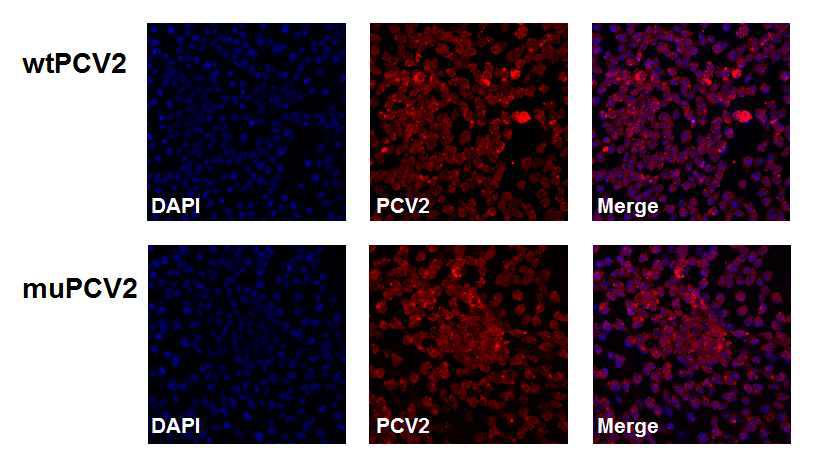
야생형 PCV2와 ORF3 돌연변이 PCV2의 PK15 세포주 감염 효율 확인 공초점현미경(confocal microscopy)으로 PCV2가 감염된 PK15 세포주에서 PCV2 ORF 2에 대한 항체로 PCV2 ORF 2가 PCV2가 감염된 PK15 세포 안에 있음을 확인하였음. DAPI, DNA를 염색; red fluorescence, PCV2 ORF 2에 대한 항체; merge, DAPI 염색과 PCV2 ORF 2에 대한 항체의 merge. PCV2 ORF 2는 PCV2의 capsid protein이기 때문에 핵 밖에서 uncoating 됨.
표
야생형 PCV2와 ORF3 돌연변이 PCV2의 PK15 세포주 감염 효율 확인 공초점현미경(confocal microscopy)으로 PCV2가 감염된 PK15 세포주에서 PCV2 ORF 2에 대한 항체로 PCV2 ORF 2가 PCV2가 감염된 PK15 세포 안에 있음을 확인하였음. DAPI, DNA를 염색; red fluorescence, PCV2 ORF 2에 대한 항체; merge, DAPI 염색과 PCV2 ORF 2에 대한 항체의 merge. PCV2 ORF 2는 PCV2의 capsid protein이기 때문에 핵 밖에서 uncoating 됨.
 표
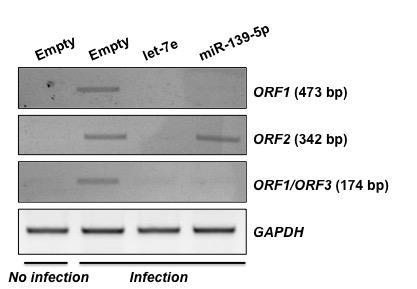
RT-PCR을 통한 let-7e 및 miR-139-5p의 PCV2 감염 저항성 확인 Empty vector, let-7e, 및 miR-139-5p를 과발현 시킨 PK15 세포에 PCV2 감염을 유도한 후 RT-PCR을 통해 각 PCV2 ORF들의 mRNA 양을 확인하여 PCV2 감염 저항성 여부를 판단함. let-7e 과발현 시 positive control (lane 2)에 비해 현저하게 PCV2 ORF들의 발현이 감소함을 확인하였으며, miR-139-5p 과발현 시 PCV2 ORF2를 제외한 ORF1, ORF3의 발현양이 감소함을 확인함.
표
RT-PCR을 통한 let-7e 및 miR-139-5p의 PCV2 감염 저항성 확인 Empty vector, let-7e, 및 miR-139-5p를 과발현 시킨 PK15 세포에 PCV2 감염을 유도한 후 RT-PCR을 통해 각 PCV2 ORF들의 mRNA 양을 확인하여 PCV2 감염 저항성 여부를 판단함. let-7e 과발현 시 positive control (lane 2)에 비해 현저하게 PCV2 ORF들의 발현이 감소함을 확인하였으며, miR-139-5p 과발현 시 PCV2 ORF2를 제외한 ORF1, ORF3의 발현양이 감소함을 확인함.
 표
숙주세포(PK15 세포) 내 PCV2의 ORF1, ORF2, ORF3 발현과 IFN-alpha, IFN-beta, IL-6의 발현량과의 상관 관계. 감염된 PK15 세포에 toll-like receptor (TLR)의 주요 ligand인 LPS (TLR2)와 poly(I:C) (TLR3)를 처리한 후, 변화하는 IFN-alpha, IFN-beta, IL-6의 발현량을 RT-PCR로 분석하였음. No, TLR ligand를 처리하지 않은 시료; LPS, LPS를 처리한 시료; Poly(I:C), poly(I:C)를 처리한 시료; mock, empty vector only.
표
숙주세포(PK15 세포) 내 PCV2의 ORF1, ORF2, ORF3 발현과 IFN-alpha, IFN-beta, IL-6의 발현량과의 상관 관계. 감염된 PK15 세포에 toll-like receptor (TLR)의 주요 ligand인 LPS (TLR2)와 poly(I:C) (TLR3)를 처리한 후, 변화하는 IFN-alpha, IFN-beta, IL-6의 발현량을 RT-PCR로 분석하였음. No, TLR ligand를 처리하지 않은 시료; LPS, LPS를 처리한 시료; Poly(I:C), poly(I:C)를 처리한 시료; mock, empty vector only.
 표
야생형 PCV2와 ORF3 돌연변이 PCV2의 PK15 세포주 감염 효율 확인 공초점현미경(confocal microscopy)으로 PCV2가 감염된 PK15 세포주에서 PCV2 ORF 2에 대한 항체로 PCV2 ORF 2가 PCV2가 감염된 PK15 세포 안에 있음을 확인하였음. DAPI, DNA를 염색; red fluorescence, PCV2 ORF 2에 대한 항체; merge, DAPI 염색과 PCV2 ORF 2에 대한 항체의 merge. PCV2 ORF 2는 PCV2의 capsid protein이기 때문에 핵 밖에서 uncoating 됨.
표
야생형 PCV2와 ORF3 돌연변이 PCV2의 PK15 세포주 감염 효율 확인 공초점현미경(confocal microscopy)으로 PCV2가 감염된 PK15 세포주에서 PCV2 ORF 2에 대한 항체로 PCV2 ORF 2가 PCV2가 감염된 PK15 세포 안에 있음을 확인하였음. DAPI, DNA를 염색; red fluorescence, PCV2 ORF 2에 대한 항체; merge, DAPI 염색과 PCV2 ORF 2에 대한 항체의 merge. PCV2 ORF 2는 PCV2의 capsid protein이기 때문에 핵 밖에서 uncoating 됨.
 표
RT-PCR을 통한 let-7e 및 miR-139-5p의 PCV2 감염 저항성 확인 Empty vector, let-7e, 및 miR-139-5p를 과발현 시킨 PK15 세포에 PCV2 감염을 유도한 후 RT-PCR을 통해 각 PCV2 ORF들의 mRNA 양을 확인하여 PCV2 감염 저항성 여부를 판단함. let-7e 과발현 시 positive control (lane 2)에 비해 현저하게 PCV2 ORF들의 발현이 감소함을 확인하였으며, miR-139-5p 과발현 시 PCV2 ORF2를 제외한 ORF1, ORF3의 발현양이 감소함을 확인함.
표
RT-PCR을 통한 let-7e 및 miR-139-5p의 PCV2 감염 저항성 확인 Empty vector, let-7e, 및 miR-139-5p를 과발현 시킨 PK15 세포에 PCV2 감염을 유도한 후 RT-PCR을 통해 각 PCV2 ORF들의 mRNA 양을 확인하여 PCV2 감염 저항성 여부를 판단함. let-7e 과발현 시 positive control (lane 2)에 비해 현저하게 PCV2 ORF들의 발현이 감소함을 확인하였으며, miR-139-5p 과발현 시 PCV2 ORF2를 제외한 ORF1, ORF3의 발현양이 감소함을 확인함.
| 과제명(ProjectTitle) : | - |
|---|---|
| 연구책임자(Manager) : | - |
| 과제기간(DetailSeriesProject) : | - |
| 총연구비 (DetailSeriesProject) : | - |
| 키워드(keyword) : | - |
| 과제수행기간(LeadAgency) : | - |
| 연구목표(Goal) : | - |
| 연구내용(Abstract) : | - |
| 기대효과(Effect) : | - |
Copyright KISTI. All Rights Reserved.

※ AI-Helper는 부적절한 답변을 할 수 있습니다.