최소 단어 이상 선택하여야 합니다.
최대 10 단어까지만 선택 가능합니다.
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
NTIS 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
DataON 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Edison 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Kafe 바로가기
| 주관연구기관 | 서울대학교 산학협력단 Seoul National University |
|---|---|
| 보고서유형 | 최종보고서 |
| 발행국가 | 대한민국 |
| 언어 | 한국어 |
| 발행년월 | 2015-03 |
| 주관부처 | 농촌진흥청 Rural Development Administration(RDA) |
| 등록번호 | TRKO201500010519 |
| DB 구축일자 | 2015-07-11 |
Ⅳ. 연구개발결과
- 토종벌 숫벌을 이용한 Whole genome sequencing을 통해 토종벌의 전체 유전체 시퀀스완성
- 이 중 화학감각 수용체 및 면역관련 유전 정보 및 pathway 분석, 사회성곤충과 비사회성곤충의 단백질 그룹 기능 비교분석
- 생리활성 펩타이드 mellitin을 동정하여 SNP 분석 및 기능분석
- 동서양 꿀벌 종의 후각 감각기능 비교 및 후각 섬모의 역할 구명
- 토종벌 조직별 long noncoding RNA 발현 분석
- SBV 감염 토종벌에 특이적으로 발현하는 m
Ⅳ. 연구개발결과
- 토종벌 숫벌을 이용한 Whole genome sequencing을 통해 토종벌의 전체 유전체 시퀀스완성
- 이 중 화학감각 수용체 및 면역관련 유전 정보 및 pathway 분석, 사회성곤충과 비사회성곤충의 단백질 그룹 기능 비교분석
- 생리활성 펩타이드 mellitin을 동정하여 SNP 분석 및 기능분석
- 동서양 꿀벌 종의 후각 감각기능 비교 및 후각 섬모의 역할 구명
- 토종벌 조직별 long noncoding RNA 발현 분석
- SBV 감염 토종벌에 특이적으로 발현하는 microRNA분석
- 국내 토종벌로부터 SBV의 분리 및 시퀀싱을 통한 바이러스 동정
- 토종벌의 전사체 NGS 시퀀싱 및 de novo assembly를 수행 및 BLAST 분석을 통한 in silico EST 라이브러리 구축
- 토종벌의 발육 단계별 유전자 발현 패턴 분석
- SBV 바이러스 감염 유충을 이용한 SBV 감염 특이 발현 유전자의 분석
- 토종벌의 내재면역계 관련 유전자의 동정 및 바이러스 감염 특이 유전자 발현 양상의 분석
- 클로닝한 SBV 유전자 절편을 이용하여 대장균에서 antiviral dsRNA의 합성
- RNAi 기법을 이용한 토종벌 유충 및 일벌의 SBV의 증식 억제 효과 검정
- 토종벌 whole body 및 독샘 cDNA library 제작으로 EST 작성, 독 및 생리활성 관련 유전자 발굴함
- 각각의 유전자 전체 염기서열을 밝히고 유전자 은행에 정보를 등록함
- 기존에 보고된 아미노산 서열 분석으로 구조 및 특성을 분석함
- 각각의 유전자를 발현하는 재조합 베큘로바이러스를 제작하고 곤충 세포주로부터 재조합단백질을 발현 시킴
- 분리한 재조합 단백질을 분리 및 정제하여 기능을 밝힘
- AcCI: chymotrypsin inhibitor, 엘라스타아제 저해활성을 박힘
- AcKTSPI: 미생물 세린 프로타아제(subtilinsin A, Protease K)에 대한 저해활성으로 항미생물 효과를 나타냄
- AcAcph-1: venom acid phosphatase의 온도 45℃, pH 4.8에서 높은 활성을 확인함
- AcICK: inhibitor cysteine knot 펩타이드의 독샘에서 살충성을 나타내는 독성분임을 밝
- AcNPC2a: Niemann-Pick disease C2의 면역 작용 (미생물 세포벽 결합)에 관여하는 사실을 밝힘
- AcICK: inhibitor cysteine knot 펩타이드의 표피 및 지방체에서 항진균 활성을 나타내며면역작용에 관여함을 밝힘
- AcApoLp: apolipophorin 유전자 서열을 분석, 구조 분석을 실시하고 재조합 단백질을 분리하여 면역작용(항균-그람양성 및 음성균-활성)에 관여함을 밝힘
- 유전변이 분석을 위한 타 꿀벌종(Apis florea)의 mitochondrial genome 분석
- 토종벌 지역변이 시료 184개체에 대한 유전분석 완료:
- 총 10개의 haplotype 확인
- 상당히 낮은 유전적 변이 가운데 기존 보고되지 않은 새로운 다수 haplotype확인
- Apis florea, A cerana 및 A. mellifera 등의 완전 mitochondrial genome의 비교분석 결과
mitochondrial DNA내 타 고변이 가능 영역 선발 : 3개 non-coding region의 분석 결과
tRNA-Met과 tRNA-Gln 사이 약 231 bp 영역 선발
- A. andreniformis의 완전 mitogenome 분석을 통해 A. andreniformis는 기존 보고된 다른 Apis 종과 달리 총 17,329 bp로 다소 크기가 크며 tRNA-Leu(CUN)이 총 5개가 존재하여 특이한 구조를 보유하고 있는 것을 확인
○ 개발 mtDNA 마커(Non-coding region 1 영역)를 이용한 국내외 집단(개체)에대한 확장 분석
- 국내외 동양종꿀벌 총 184개체에 대한 분석
- 총 9개의 haplotype을 얻었으며 이들의 길이는 230~231 bp
- 전년도 연구된 범용 mtDNA 마커와 비교시 당해연도의 NC1이 보다 변별력을 가진 마커로서 활용 가능
- 국내외 집단에 대한 NC1 분석결과, 분석된 동양종꿀벌은 크게 두 개의 그룹 (Group A,
B)으로 나누어지는 것을 확인함 (p > 0,05%): Group A는 우리나라의 청송군, 양구군, 평창군, 그리고 중국의 Changbai Mountain, Kunming 등 6개 지역으로 구성되어 있음
- Group B는 나머지 지역인 우리나라 청주시, 남원시, 이천시, 청원군 및 중국의 베이징으로구성되어 있음
○ 핵 지놈 영역 마커 탐색 (2개 이상)
- 엑손 사이에 위치하는 인트론 영역 6개 선발을 선발하여 스크리닝한 결과 2개 영역을 제외한 4개의 영역이 변별력을 가지고 있는 것으로 판단되었음
○ 핵 지놈 영역 마커를 이용한 국내외 개체에 대한 확장 분석
- 국내 9개 지점 및 중국, 베트남, 태국 등 국외 7개 지점 (총 16개 지역)에서 채집된 184개체에 대한 3개 이상의 클론분석 완료
- MRJP9 gene 인터론 영역 분석 결과, 307~322 bp 얻음으며 총 552개 콜론 중 222개의 sequence type을 확인하였음 (ACMR01~ACMR222)
- White gene 인터론 영역 분석 결과, 157~163 bp 얻음 총 552개 콜론 중 141개의 sequence type을 확인하였음 (ACWH01~ACWH141)
The honey bee has long been used for a pollinator as well as honey production in human history. There are two major honey bee species in the world, Apis mellifera and Apis cerana. Since 2003, world has been experiencing the critical decrease of honey bee colony size due to unconfirmed reasons, which
The honey bee has long been used for a pollinator as well as honey production in human history. There are two major honey bee species in the world, Apis mellifera and Apis cerana. Since 2003, world has been experiencing the critical decrease of honey bee colony size due to unconfirmed reasons, which is called colony collapse disorder (CCD). Also, as a counterpart, in most southern and eastern parts of Asia were suffering from pivotal virus disease in A. cerana, which is called sacbrood virus infection. A. cerana is a major pollinator in most Asian countries for thousands years. By year of 2012, about 90% of Apis cerana were dead due to this disease in Korea.
Since 2006 when genomics information of A. mellifera was announced, many studies on the molecular and neurobiological mechanisms of bee biology have been carried out. Recently, with huge development of bioinformatics and sequencing techniques, it has been important to identify and characterize the functional genes responsible for social behaviors, immune system, defense, and caste development. Therefore, in this project we have been focusing on the characterization of novel genes associated with immunity and chemosensory reception as well as nutrition. In social insects like honey bees, chemical communication plays a critical role in colony maintenance, development, behavioral regulation, and caste transition, as well as in defense behaviors. It has been reported that A. cerana is more efficient to remove ectoparasitic mites such as Varroa distructor mainly based on olfactory detection abilities, compared to A. mellifera.
In order to understand the differences of olfactory characteristics in the two species, we compared the distribution of sensory hairs on the antennae and antennal olfactory responses, using electron microscopy, electrophysiological recording and molecular expression level of odorant receptors. Our study demonstrated that the antennae of A. cerana have more olfactory sensilla than A. mellifera, responding more strongly to various floral volatile compounds. At the molecular level, olfactory co-receptor (Orco), which makes heterodimers with other conventional olfactory receptors, is more abundantly expressed in the antenna of A. cerana than of A. mellifera. In addition, we identified two different forms of insect-derived antimicrobial peptides (AMPs) exist in A. cerana and A. mellifera, which showed different effects on antimicrobial properties and pharmacological activities such as anti-inflammation and anticancer properties. We found that single nucleotide polymorphism (SNP) occurring in natural melittin gene of honey bee populations, which is one of the potent AMP in bee venoms. We found that the novel SNP of melittin gene exists in these two honey bee species, A. mellifera and A. cerana.Nine polymorphisms were identified within the coding region of the melittin gene, of which one polymorphism that resulted inserine (Ser) to asparagine (Asp) substitution that can potentially effect on biological activities of melittin peptide. Finally, using hiseq2000 and other new RNA sequencing techniques, we identified immune genes and chemosensory genes, neuropeptide genes in A. cerana with expression patterns in different parts of body tissues. Taken together, these novel genes play a role in interaction with many stress factors and disease pathogens, which will pave the way for future directions for beeding program in A. cerana.
The Asian honeybee, Apis cerana, is a native honeybee species in Korea which is important in agriculture for pollination and honey production. Since 2009, A. cerana in Korea is endangered because of the outbreak of sacbrood virus (SBV) disease which caused over 75% of coloy loss. To study the molecular biology of the honeybee virus disease, SBV was isolated and identified from local samples, and virus replication pattern was investigated with qPCR, and functional genomics approach was employed with next generation sequencing technique.
For better understanding of the physiology of A. cerana, high-throughput Illumina transcriptome sequencing was performed to analyze the gene expression profiles of queen, worker, and larva. A total of 219,799,682 clean reads corresponding to 22.2 Gb of nucleotide sequences was obtained from the whole body total RNA samples. The Apis mellifera reference mRNA sequence database was used to measure the gene expression level with Bowtie2 and eXpress software, and the Illumina short reads were then mapped to 11,459 out of 11,736 A. mellifera reference genes. Total of 9,221 genes with FPKM value greater than 5 of each sample group were subjected to eggNOG with BLASTX for gene ontology analysis. The differential gene expression between queen and worker, and worker and larva were analyzed to screen the overexpressed genes in each sample group. In the queen and worker sample group, total of 1,766 genes were differentially expressed with 887 and 879 genes overexpressed over two folds in queen and worker, respectively. In the worker and larva sample group, total of 1,410 genes were differentially expressed with 1,009 and 401 genes overexpressed over two folds in worker and larva, respectively.
Also an in silico EST library of A. cerana was constructed with 101 base pair-end illumine sequencing result of the transcriptome of A. cerana larva. A total of 205,597,180 and 189,276,088 of clean reads corresponding to 20.8 Gb and 19.1 Gb from SBV carrying and control A. cerana larva, respectively. By using Trinity de novo assembly software, total of 67,750 contigs were constructed, and 21,931 of them were annotated with BLASTX and eggNOG software.
The gene expression pattern of SBV carrying A. cerana larva was analyzed with the same procedure previously described. The top 20,000 of the most frequently mapped transcripts were subjected to Bowtie2 and eXpress software to show that 315 and 2,263 of the annotated transcripts were up- and down-regulated, respectively, while the expression level of ribosomal protein genes which are supposed not to be affected by virus infection were about same between the experimental and control sample. Especially the 109 genes belonging to 14 of gene families of innate immune pathway showed differential gene expression upon virus infection.
To investigate the RNAi-silencing of SBV, A. cerana larvae and workers were fed with antiviral dsRNA. The partial fragment of SBV VP1 coat protein and RNA dependent RNA polymerase were cloned into the RNA transcription vector, L4440, and the dsRNA was extracted from the transformed HT115 E. coli. In this study, the suppression of virus was measured by qPCR to show that the antiviral dsRNA may suppress SBV level up to 9.6 times in both worker bees, and the suppression level was dependent to the amount of dsRNA inoculum.
The honeybee is a social insect species and an economically important insect that plays an essential role in the pollination of flowering plants (Velthuis and Doorn, 2006) and the supply of several products, such as honey, royal jelly, propolis, pollen, wax, and venom (Cherniack, 2010). Honeybee-derived products have been used as a staple of traditional medicine for centuries in Asian countries such as Korea, Japan, and China (Cherniack, 2010).
Bee venom is composed of a variety of enzymes, peptides, and biogenic amines (Hoffman, 2006; Hoffman and Jacobson, 1996; Peiren et al., 2005; Son et al., 2007; Winningham et al., 2004). Some bee venom components can cause life-threatening allergic reactions due to an immediate hypersensitivity-induced reaction that leads to anaphylaxis (Fitzgerald and Flood, 2006; Golden, 2007). Honeybee venom allergens are as follows: Api m 1 (phospholipase A2), Api m 2 (hyaluronidase), Api m 3 (acid phosphatase), Api m 4 (melittin), Api m 5 (dipeptidylpeptidase IV), Api m 6, Api m 7 (CUB serine protease), Api m 8 (carboxylesterase), and Api m 9 (serine carboxypeptidase). Two major components of honeybee venom are melittin and phospholipase A2 (PLA2) (Gauldie et al., 1976; Six and Dennis, 2000) and are the most-studied bee venom components because of their anticancer effects (Heinen and da Veiga, 2011).In addition to melittin and PLA2, other bee venom components have been studied as promising sources of bioactive compounds. Our previous studies demonstrated a novel mechanism by which bumblebee venom affects the hemostatic system via venom serine protease inhibitor-mediated antifibrinolytic activity as well as via venom serine protease-mediated fibrin(ogen)olytic activities (Choo et al., 2010, 2012; Qiu et al., 2011, 2013). Bee venom is considered a rich source of pharmacologically active components, and venom components have been intensively studied as potential compounds on which to base novel pharmaceuticals (Heinen and da Veiga, 2011; Son et al., 2007). The protection of bees as pollinators of flowering plants (Velthuis and Doorn, 2006) and the utilization of bees as suppliers of natural products have been subjects of worldwide focus (Cherniack, 2010). However, although the importance of honeybee, the function of Korean honeybee-derived venom components and products has not been described until now.
Here, we report the molecular characterization of the Korean honeybee (Apis cerana) chymotrypsin inhibitor (AcCI), venom-derived Kazal-type serine protease inhibitor (AcKTSPI), venom acid phosphatase Acph-1-like protein (AcAcph-1), Niemann-Pick disease type C2 (AcNPC2a), inhibitor cysteine knot (AcICK) peptide, and apolipophorin III (apoLp-III).The final goal of the present work is to study on venom-derived genes of Korean honeybee, Apis cerana and its functional analysis. This work is based on the following content and scope:
1) Screening and expression of venom-derived genes from Korean honeybee
2) Functional analysis of venom-derived genes from Korean honeybee
3) Screening and functional analysis of Korean honeybee-derived bioactive peptide
Study on venom-derived genes of Korean bee, Apis cerana and its functional analysis
1) Anti-elastolytic activity of a honeybee (Apis cerana) chymotrypsin inhibitor
The honeybee is an important insect species in global ecology, agriculture, and alternative medicine. While chymotrypsin and trypsin inhibitors from bees show activity against cathepsin G and plasmin, respectively, no anti-elastolytic role for these inhibitors has been elucidated. In this study, we identified an Asiatic honeybee (Apis cerana) chymotrypsin inhibitor (AcCI), which was shown to also act as an elastase inhibitor. AcCI was found to consist of a 65-amino acid mature peptide that displays ten cysteine residues.When expressed in baculovirus-infected insect cells, recombinant AcCI demonstrated inhibitory activity against chymotrypsin (Ki 11.27 nM), but not trypsin, defining a role for AcCI as a honeybee-derived chymotrypsin inhibitor. Additionally, AcCI showed no detectable inhibitory effects on factor Xa, thrombin, plasmin, or tissue plasminogen activator; however, AcCI inhibited human neutrophil elastase (Ki 61.05 nM), indicating that it acts as an anti-elastolytic factor. These findings constitute molecular evidence that AcCI acts as a chymotrypsin/elastase inhibitor.
2) Antimicrobial activity of a honeybee (Apis cerana) venom Kazal-type serine protease inhibitor
Insect-derived Kazal-type serine protease inhibitors exhibit thrombin, elastase, plasmin, proteinase K, or subtilisin A inhibition activity, but so far, no functional roles for bee-derived Kazal-type serine protease inhibitors have been identified. In this study, a bee(Apis cerana) venom Kazal-type serine protease inhibitor (AcKTSPI) that acts as a microbial serine protease inhibitor was identified. AcKTSPI contained a single Kazal domain that displayed six conserved cysteine residues and a P1 threonine residue. AcKTSPI was expressed in the venom gland and was present as a 10-kDa peptide in bee venom. Recombinant AcKTSPI Kazal domain (AcKTSPI-Kd) expressed in baculovirus-infected insect cells demonstrated inhibitory activity against subtilisin A (Ki67.03 nM) and proteinase K (Ki 91.53 nM), but not against α-chymotrypsin or trypsin,which implies a role for AcKTSPI as a microbial serine protease inhibitor. However,AcKTSPI-Kd exhibited no detectable inhibitory effects on factor Xa, thrombin, tissue plasminogen activator, or elastase. Additionally, AcKTSPI-Kd bound directly to Bacillus subtilis, Bacillus thuringiensis, Beauveria bassiana, and Fusarium graminearum but not to Escherichia coli . Consistent with these findings, AcKTSPI-Kd showed antibacterial activity against Gram-positive bacteria and antifungal activity against both plant-pathogenic and entomopathogenic fungi. These findings constitute molecular evidence that AcKTSPI acts as an inhibitor of microbial serine proteases. This paper provides a novel view of the antimicrobial functions of a bee venom Kazal-type serine protease inhibitor.
3) Molecular characterization of a venom acid phosphatase Acph-1-like protein from the Asiatic honeybee Apis cerana
Bee venom contains a variety of peptides and enzymes, including acid phosphatases. An acid phosphatase has been identified from European honeybee (Apis mellifera) venom. However, although the amino acid sequence is known, no functional information is currently available for bee venom acid phosphatase Acph-1-like proteins. In this study, an Asiatic honeybee (Apis cerana) venom acid phosphatase Acph-1-like protein (AcAcph-1) was identified. The analysis of the predicted AcAcph-1 amino acid sequence revealed high levels of identity with other bee venom acid phosphatase Acph-1-like proteins. Recombinant AcAcph-1 was expressed as a 64-kDa protein in baculovirus-infected insect cells. The enzymatic properties of recombinant AcAcph-1, determined using p-nitrophenyl phosphate (p-NPP) as a substrate, showed the highest activity at 45 ℃ and pH 4.8. Northern and western blot analyses showed that AcAcph-1 was expressed in the venom gland and was present as a 64-kDa protein in bee venom. In addition, N-glycosylation of AcAcph-1 was revealed by tunicamycin treatment of recombinant virus-infected insect Sf9 cells and by glycoprotein staining of purified recombinant AcAcph-1. Our findings show that AcAcph-1 functions as a venom acid phosphatase. This paper provides the first evidence of the role of a bee venom acid phosphatase Acph-1-like protein.
4) Molecular characterization of a Niemann-Pick disease type C2 protein from the honeybee Apis cerana
Drosophila Niemann–Pick disease type C2 (NPC2) proteins play roles in sterol homeostasis, steroid biosynthesis, and innate immune signaling pathways. In this study, a bee (Apis cerana) NPC2a protein (AcNPC2a) that might function in innate immune reactions was identified. AcNPC2a consisted of 148 amino acids, which included six conserved cysteine residues. Recombinant AcNPC2a protein (expressed in baculovirus-infected insect cells) bound directly to live Escherichia coli , Bacillus thuringiensis, and Beauveria bassiana; however, AcNPC2a did not show antimicrobial activity against these microorganisms. Nevertheless, the expression of AcNPC2a was significantly induced in the fat body of A. cerana worker bees after injection with E. coli ,B. thuringiensis, or B. bassiana. Our data suggest a role for AcNPC2a in innate immunity that is induced in response to microbial challenge and binds directly to the cell walls of bacteria and fungi. These findings provide insight into the role of AcNPC2a during the innate immune response following bacterial and fungal infection.
5) Dual function of a bee (Apis cerana) inhibitor cysteine knot peptide that acts as an antifungal peptide and insecticidal venom toxin
Inhibitor cysteine knot (ICK) peptides exhibit ion channel blocking, insecticidal, and antimicrobial activities, but currently, no functional roles for bee-derived ICK peptides have been identified. In this study, a bee (Apis cerana) ICK peptide (AcICK) that acts as an antifungal peptide and as an insecticidal venom toxin was identified. AcICK contains an ICK fold that is expressed in the epidermis, fat body, or venom gland and is present as a 6.6-kDa peptide in bee venom. Recombinant AcICK peptide (expressed in baculovirus-infected insect cells) bound directly to Beauveria bassiana and Fusarium graminearum, but not to Escherichia coli or Bacillus thuringiensis. Consistent with these findings, AcICK showed antifungal activity, indicating that AcICK acts as an antifungal peptide. Furthermore, AcICK expression is induced in the fat body and epidermis after injection with B. bassiana. These results provide insight into the role of AcICK during the innate immune response following fungal infection. Additionally, we show that AcICK has insecticidal activity. Our results demonstrate a functional role for AcICK in bees: AcICK acts as an antifungal peptide in innate immune reactions in the body and as an insecticidal toxin in venom. The finding that the AcICK peptide functions with different mechanisms of action in the body and in venom highlights the two-pronged strategy that is possible with the bee ICK peptide.
6) Apolipophorin III from honeybees (Apis cerana) exhibits antibacterial activity
Apolipophorin III (apoLp-III) is involved in lipid transport and innate immunity in insects. In this study, an apoLp-III protein that exhibits antibacterial activity was identified in honeybees (Apis cerana). A. cerana apoLp-III cDNA encodes a 193 amino acid sequence that shares high identity with other members of the hymenopteran insect apoLp-III family. A. cerana apoLp-III is expressed constitutively in the fat body, epidermis, and venom gland and is detected as a 23-kDa protein. A. cerana apoLp-III expression is induced in the fat body after injection with Escherichia coli , Bacillus thuringiensis, or Beauveria bassiana.However, recombinant A. cerana apoLp-III (expressed in baculovirus-infected insect cells) binds directly to E. coli and B. thuringiensis but not to B. bassiana. Consistent with these findings, A. cerana apoLp-III exhibited antibacterial activity against both Gram-negative and Gram-positive bacteria. These results provide insight into the role of A.cerana apoLp-III during the innate immune response following bacterial infection. The study of molecular biological aspects of A. cerana is fundamentally important for eventual breeding and distribution of new breeds. Further. accumulation of a diverse level of genetic information. development of new molecular marker and genomic illustration have essential importance for value enhancement and brand making of invaluable domestic genetic resources.
Therefore, in this research, we investigated several levels of molecular genetic aspects. First of all, current progress on molecular genetic studies was investigated to select proper molecular markers for the investigation of genetic diversity of Korean A. cerana via domestic and foreign literature search and selected a universally utilized molecular marker, the non-coding region of mitochondrial genome that is located between tRNALeuand COII(termed NC2). Using the marker, a total of 184 bees collected from nine Korean localities and seven overseas localities (five from China, one from Vietnam, and one from Thailand) were sequenced and compared with ~500 worldwide sequence information to understand phylogenetic relationships of Korean A. cerana and worldwide samples. Ten haplotypes were found from NC2: four undiscovered haplotypes were found in Korea and China,respectively, whereas previously reported haplotypes were found in Vietnam (Japan1) and Thailand (Japan1 and IndiaB4).
A phylogenetic analysis confirmed that Korean A. cerana belonged to the Mainland Asian group. The most widespread haplotype in Mainland Asia was Japan1, and it was also found dominantly in samples from other countries including Korea, accounting for 71%. Such dominance of Japan1 suggests extensive gene flow onto Mainland Asia mediated by Japan1.
However, the marker provide a limited resolution to infer Korean populations of A. cerana. Thus, additional marker was required to better understand the genetic diversity and relationships of Korean A. cerana. Therefore, we sequenced the full-length mitochondrial genome of other Apis species such as A. dorsata and A. andreniformis. These sequences were aligned with A. cerana full-length mitochondrial genome and found three additional non-coding region in the A. cerana. Among them one non-coding region located between tRNAMetand tRNAGln in mitochondrial genome of A. cerana (termed NC1) turned out to be useful for further investigation and used for phylogenetic relationships in Korean A. cerana.An NC1-based phylogenetic analysis revealed the presence of two phylogenetic groups in Korea, without any region-based clustering. These results suggest that A. cerana in Korea may have been introduced to Korea from two different sources in India via China long ago. Characteristic A. cerana beekeeping, such as continuous hives in mountainous locations, may have allowed A. cerana stocks to remain as they were introduced but more likely human intervention along with natural dispersal may have facilitated mingling of random populations.
Also, we selected two nuclear intron markers (MRJP9 and White gene intron) after prior investigation on PCR availability and variability of six-region sequences. Sequencing of each three clones from all 184 samples provided a total of 222 sequence types for MRJP9 and 141 sequence types for White gene intron. Further, these intron sequences possessed a proper level of percent sequence diversity and G/C content enough to utilize for further investigation of genetic diversity of Korean A. cerana. For the perspective of diversity these markers also can further be utilized for the diversity investigation. Possibly due to evolutionary difference of the introns in speed the consequent population relationships were not identical. However, the two intron results indicates that the introns are promising for further study of A. cerana populations and Koran A. cerana is overall composed of more than two different origin.
 표
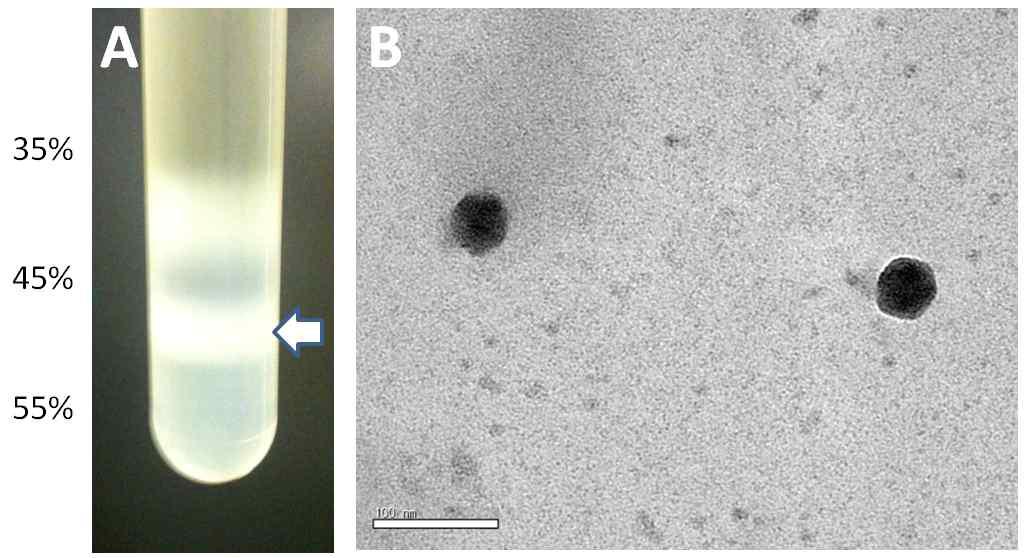
Sucrose gradient ultracentrifuge purification of SBV. A. A crude virus extract from A. cerana was subjected to a sucrose gradient of 35%~55% for ultracentirifugation. The virus band formed between 45% and 55% layer was collected after ultracentrifugation with Beckman SW40.1 rotor for 1 hr at 30,000rpm. B. SBV particles purified by ultracentrifugation was stained with 0.5% uranyl acetate and observed by transmission electron microscope. The sizebar indicates 100nm.
표
Sucrose gradient ultracentrifuge purification of SBV. A. A crude virus extract from A. cerana was subjected to a sucrose gradient of 35%~55% for ultracentirifugation. The virus band formed between 45% and 55% layer was collected after ultracentrifugation with Beckman SW40.1 rotor for 1 hr at 30,000rpm. B. SBV particles purified by ultracentrifugation was stained with 0.5% uranyl acetate and observed by transmission electron microscope. The sizebar indicates 100nm.
 표
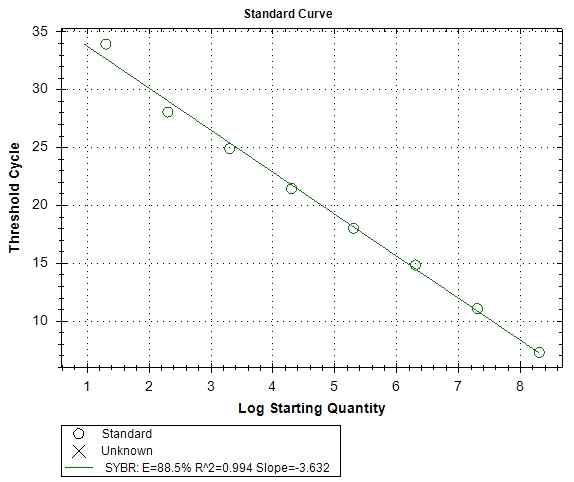
Standard curve of the SBV RdRp specific primer sets. A plasmid containing 0.8kb fragment of SBV RdRp was used for dilution series of PCR template. 1x108 plasmid molecules were diluted over a 8-log scale down to 10 molecules. The PCR efficiency, R2, and slope were 88.5%, 0.99, and –3.6, respectively.
표
Standard curve of the SBV RdRp specific primer sets. A plasmid containing 0.8kb fragment of SBV RdRp was used for dilution series of PCR template. 1x108 plasmid molecules were diluted over a 8-log scale down to 10 molecules. The PCR efficiency, R2, and slope were 88.5%, 0.99, and –3.6, respectively.
 표
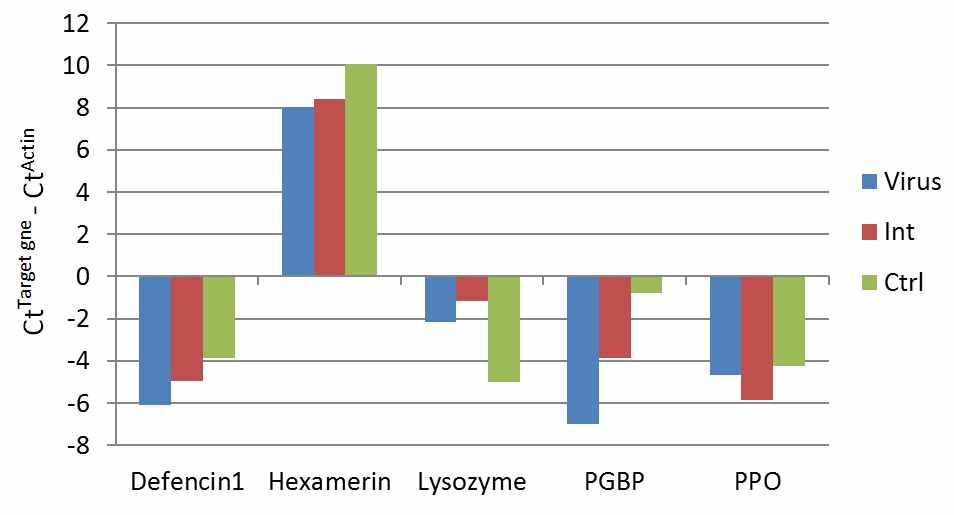
The SBV concentration-dependent gene expression patterns of invertebrate immune related genes. The X-axis represents the concentration of target genes in each sample by 2(Ct value of target gene – Ct value of ctin gene). Virus; A. cerana carrying h high amount of SBV, Int; A. cerana carrying intermediate amount of SBV, Ctrl; negative control. Based on qPCR study, the Virus carried four times more SBV than Int does.
표
The SBV concentration-dependent gene expression patterns of invertebrate immune related genes. The X-axis represents the concentration of target genes in each sample by 2(Ct value of target gene – Ct value of ctin gene). Virus; A. cerana carrying h high amount of SBV, Int; A. cerana carrying intermediate amount of SBV, Ctrl; negative control. Based on qPCR study, the Virus carried four times more SBV than Int does.
 표
Alignment of the amino acid sequences for AcAcph-1 and known bee venom acid phosphatase Acph-1-like proteins. A vertical arrow indicates the end of the signal peptide. Potential N-glycosylation sites are indicated by crosses. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ710421), Apis mellifera (GenBank accession no. XP_006569977), Bombus impatiens (GenBank accession no. XP_003486582), and Megachile rotundata (GenBank accession no. XP_003706109). The AcAcph-1 sequence was used as the reference for the identity/similarity (Id/Si) values.
표
Alignment of the amino acid sequences for AcAcph-1 and known bee venom acid phosphatase Acph-1-like proteins. A vertical arrow indicates the end of the signal peptide. Potential N-glycosylation sites are indicated by crosses. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ710421), Apis mellifera (GenBank accession no. XP_006569977), Bombus impatiens (GenBank accession no. XP_003486582), and Megachile rotundata (GenBank accession no. XP_003706109). The AcAcph-1 sequence was used as the reference for the identity/similarity (Id/Si) values.
 표
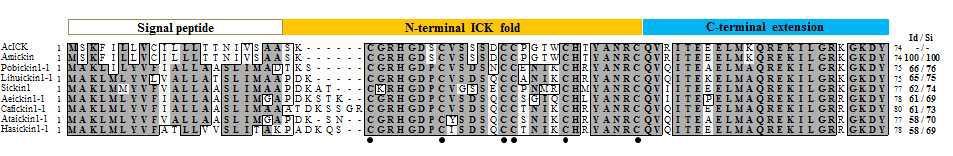
The alignment of the amino acid sequences for AcICK and known ICK peptides. The characteristic cysteine residues are indicated by solid circles. The signal peptide, N-terminal ICK fold, and C-terminal extension are shown. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ530970), A. mellifera (GenBank accession no. XP_006560077), Pogonomyrmex barbatus (GenBank accession no.ADIH01011549), Linepithema humile (GenBank accession no. ADOQ01003051), Solenopsis invicta (GenBank accession no. JQ700385), Acromyrmex echinatior (GenBank accession no.AEVX01007421), Camponotus floridanus (GenBank accession no. AEAB01015096), Atta cephalotes (GenBank accession no. ADTU01022195), and Harpegnathos saltator (GenBank accession no. AEAC01004275). The AcICK sequence was used as the reference for the identity/similarity (Id/Si) values.
표
The alignment of the amino acid sequences for AcICK and known ICK peptides. The characteristic cysteine residues are indicated by solid circles. The signal peptide, N-terminal ICK fold, and C-terminal extension are shown. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ530970), A. mellifera (GenBank accession no. XP_006560077), Pogonomyrmex barbatus (GenBank accession no.ADIH01011549), Linepithema humile (GenBank accession no. ADOQ01003051), Solenopsis invicta (GenBank accession no. JQ700385), Acromyrmex echinatior (GenBank accession no.AEVX01007421), Camponotus floridanus (GenBank accession no. AEAB01015096), Atta cephalotes (GenBank accession no. ADTU01022195), and Harpegnathos saltator (GenBank accession no. AEAC01004275). The AcICK sequence was used as the reference for the identity/similarity (Id/Si) values.
 표
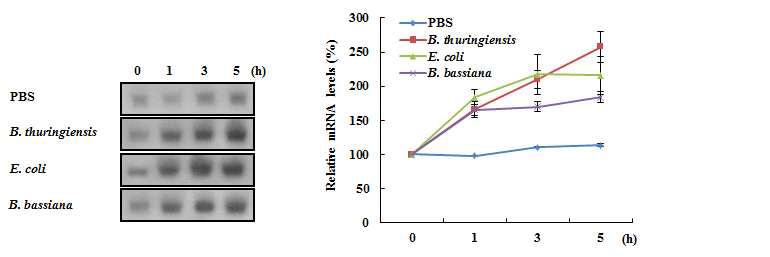
Transcriptional expression profiles of the A. cerana apoLp-III gene in the fat body of A. cerana worker bees induced by E. coli , B. thuringiensis, or B. bassiana injection. A cerana worker bees injected with PBS were used as injection controls. Total RNA was isolated from the fat body of worker bees at different time points as indicated above the corresponding lanes. A. cerana apoLp-I I I transcripts were analyzed using northern hybridization with radiolabeled A. cerana apoLp-I I I cDNA (upper). The levels of A. cerana apoLp-I I I mRNA are reported as the means of three measurements that were calculated relative to the levels at 0 h before PBS or microbial injection (shown as 100%) (lower). Bars represent the means ± SD.
표
Transcriptional expression profiles of the A. cerana apoLp-III gene in the fat body of A. cerana worker bees induced by E. coli , B. thuringiensis, or B. bassiana injection. A cerana worker bees injected with PBS were used as injection controls. Total RNA was isolated from the fat body of worker bees at different time points as indicated above the corresponding lanes. A. cerana apoLp-I I I transcripts were analyzed using northern hybridization with radiolabeled A. cerana apoLp-I I I cDNA (upper). The levels of A. cerana apoLp-I I I mRNA are reported as the means of three measurements that were calculated relative to the levels at 0 h before PBS or microbial injection (shown as 100%) (lower). Bars represent the means ± SD.
 표
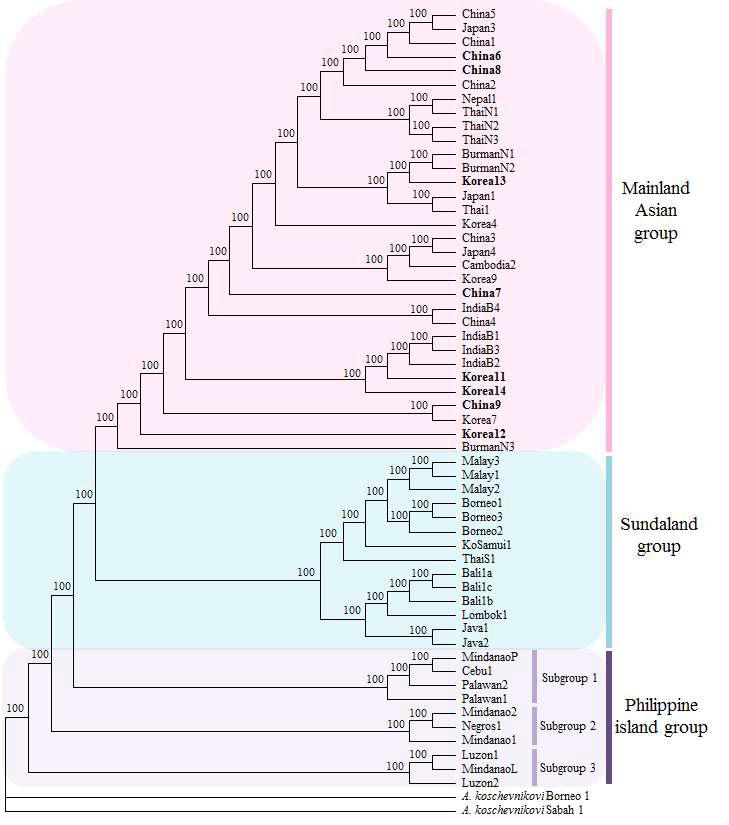
Relationships among the Apis cerana NC2 haplotypes. The tree was constructed using the minimumevolution method that was rooted using two haplotypes of A. koschevnikovi (GenBank accession numbers AB072437 and AB072438; Takahashi et al. 2002) and presented as 50% majority-rule consensus tree. Eight haplotypes were found new in this study (bold), and 48 haplotypes reported previously were included in the analysis. Nodal support was obtained by 1,000 replicates.
표
Relationships among the Apis cerana NC2 haplotypes. The tree was constructed using the minimumevolution method that was rooted using two haplotypes of A. koschevnikovi (GenBank accession numbers AB072437 and AB072438; Takahashi et al. 2002) and presented as 50% majority-rule consensus tree. Eight haplotypes were found new in this study (bold), and 48 haplotypes reported previously were included in the analysis. Nodal support was obtained by 1,000 replicates.
 표
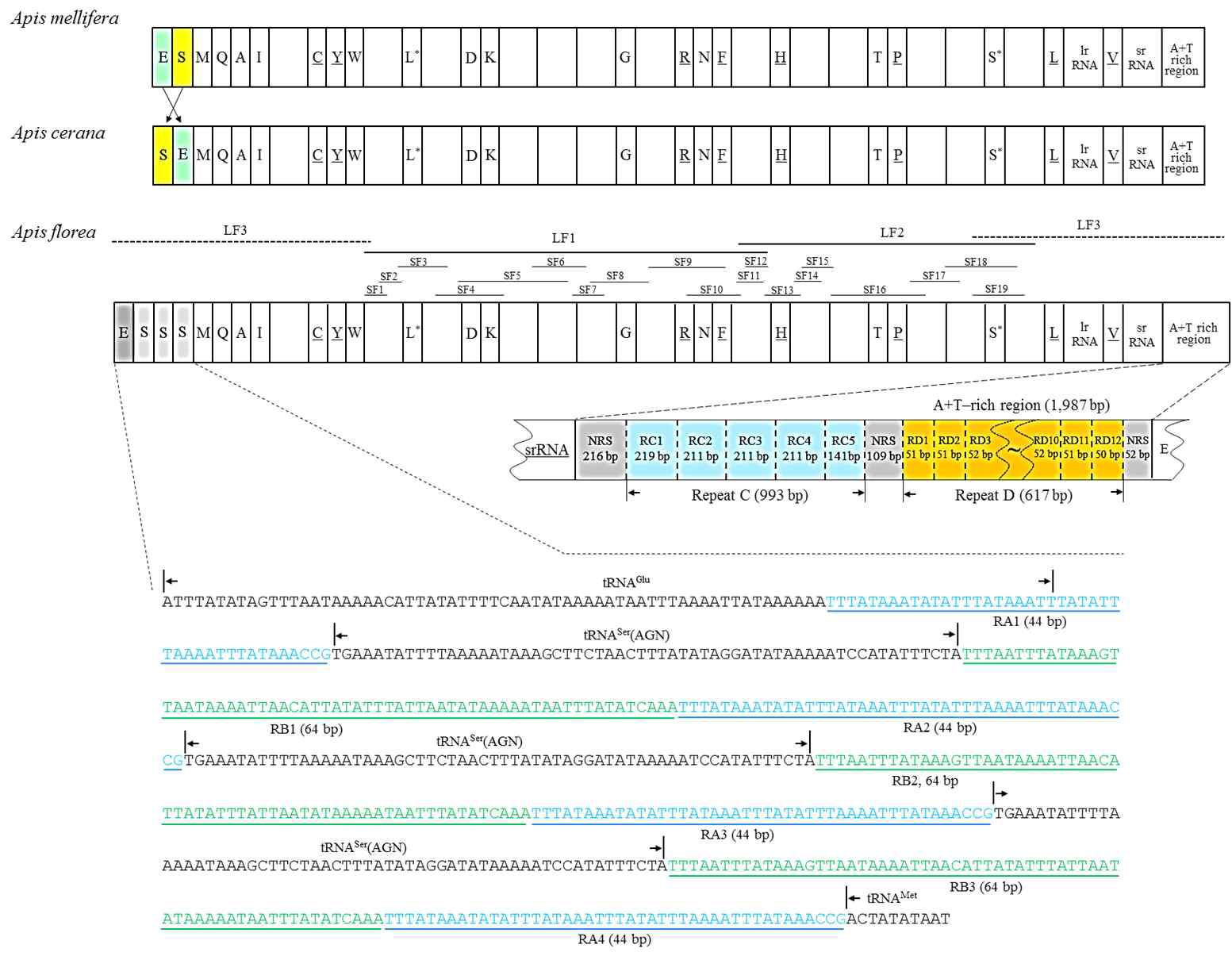
Linear map of Apis florea mitochondrial genome, with the schematic map of triplicated tRNA-Ser(AGN) regionlocated between tRNA-Glu and tRNA-Met and the A+T-rich region. The blank boxes are protein-coding genes that are identical in all hymenopteran family Apidae in arrangement. tRNAs are denoted as one-letter symbols in accordance with the IUPAC-IUB single-letter amino acid codes. The one-letter symbols L, L*, S and S* denote tRNA-Leu(CUN), tRNA-Leu(UUR), tRNA-Ser(AGN), and tRNA-Ser(UCN), respectively. Gene names that are not underlined indicate a clockwise direction of transcription, whereas underlines indicate a counter-clockwise direction. Sequences of triplicated tRNAser(AGN) region are shown with the precedent (RA1 ~ RA3) and following repeats (RB1 ~RB3) plus an extra RA4. SF and LF refer short and long fragments, respectively. LF3 was sequenced by shotgun method, whereas LF1 and LF2 were used as temperate for the amplification and sequencing of the 19 SFs. Schematic map of 1,987-bp long A+T-rich region is composed of two repeat units, named as RC (RC1 ~ RC4) and RD (RD1 ~ RD12) encompassed by three non-repeat sequence (NRS). RD4 ~ RD9 within dotted wavy line were omitted in this map.
표
Linear map of Apis florea mitochondrial genome, with the schematic map of triplicated tRNA-Ser(AGN) regionlocated between tRNA-Glu and tRNA-Met and the A+T-rich region. The blank boxes are protein-coding genes that are identical in all hymenopteran family Apidae in arrangement. tRNAs are denoted as one-letter symbols in accordance with the IUPAC-IUB single-letter amino acid codes. The one-letter symbols L, L*, S and S* denote tRNA-Leu(CUN), tRNA-Leu(UUR), tRNA-Ser(AGN), and tRNA-Ser(UCN), respectively. Gene names that are not underlined indicate a clockwise direction of transcription, whereas underlines indicate a counter-clockwise direction. Sequences of triplicated tRNAser(AGN) region are shown with the precedent (RA1 ~ RA3) and following repeats (RB1 ~RB3) plus an extra RA4. SF and LF refer short and long fragments, respectively. LF3 was sequenced by shotgun method, whereas LF1 and LF2 were used as temperate for the amplification and sequencing of the 19 SFs. Schematic map of 1,987-bp long A+T-rich region is composed of two repeat units, named as RC (RC1 ~ RC4) and RD (RD1 ~ RD12) encompassed by three non-repeat sequence (NRS). RD4 ~ RD9 within dotted wavy line were omitted in this map.
 표
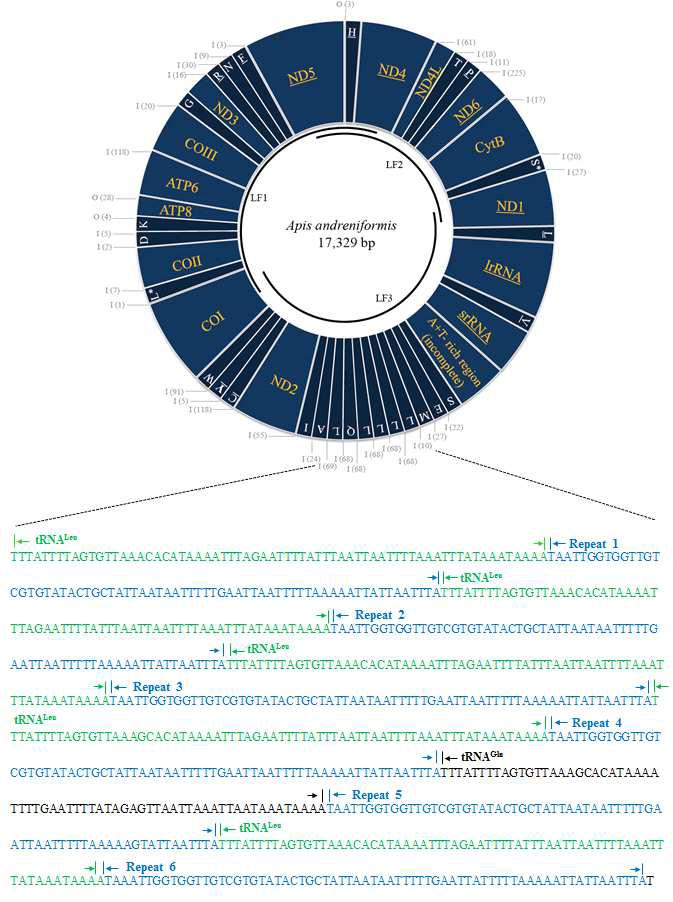
Circular map of the Apis andreniformis mitochondrial genome with the tandemly repeatedtRNA-Leu(CUN) region sequences. Four additional tRNA-Leu(CUN) were found between tRNA-Met and tRNA-Gln, and one additional tRNA-Leu(CUN) was found between tRNA-Gln and tRN-AAla. A nearly identical 68-bp long repeat sequence (repeats 1–6) abutted the 5′-end of each additional tRNA-Leu(CUN) and tRNA-Gln.
표
Circular map of the Apis andreniformis mitochondrial genome with the tandemly repeatedtRNA-Leu(CUN) region sequences. Four additional tRNA-Leu(CUN) were found between tRNA-Met and tRNA-Gln, and one additional tRNA-Leu(CUN) was found between tRNA-Gln and tRN-AAla. A nearly identical 68-bp long repeat sequence (repeats 1–6) abutted the 5′-end of each additional tRNA-Leu(CUN) and tRNA-Gln.
 표
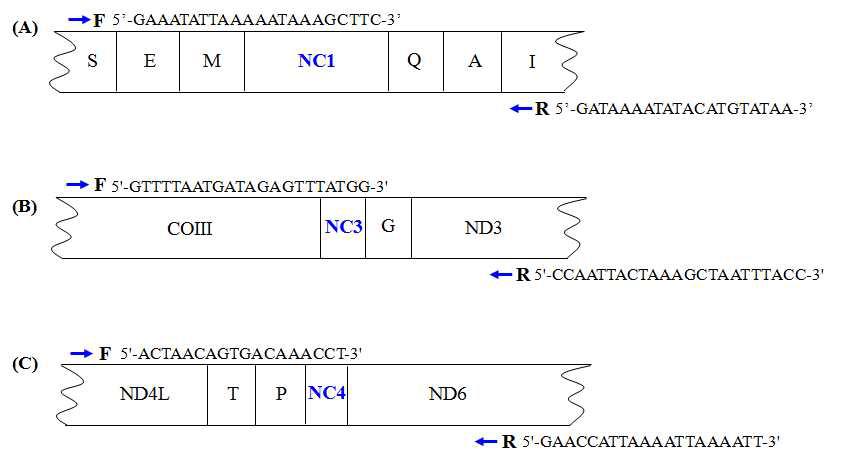
Locations of three major non-coding regions located in the A. cerana mitochondrial genome. Positions ofSer Ile(A) NC1 located between tRNA and tRNA , NC3 located between COIII, and ND3 NC4 located between ND4L and ND6. S, E, M, Q, A, I, G, T and P, respectively, indicate tRNASer, tRNAGlu, tRNAMet, tRNAGln, tRNAAla, tRNAIle, tRNAGly, tRNAThr, and tRNAPro, respectively.
표
Locations of three major non-coding regions located in the A. cerana mitochondrial genome. Positions ofSer Ile(A) NC1 located between tRNA and tRNA , NC3 located between COIII, and ND3 NC4 located between ND4L and ND6. S, E, M, Q, A, I, G, T and P, respectively, indicate tRNASer, tRNAGlu, tRNAMet, tRNAGln, tRNAAla, tRNAIle, tRNAGly, tRNAThr, and tRNAPro, respectively.
 표
Sucrose gradient ultracentrifuge purification of SBV. A. A crude virus extract from A. cerana was subjected to a sucrose gradient of 35%~55% for ultracentirifugation. The virus band formed between 45% and 55% layer was collected after ultracentrifugation with Beckman SW40.1 rotor for 1 hr at 30,000rpm. B. SBV particles purified by ultracentrifugation was stained with 0.5% uranyl acetate and observed by transmission electron microscope. The sizebar indicates 100nm.
표
Sucrose gradient ultracentrifuge purification of SBV. A. A crude virus extract from A. cerana was subjected to a sucrose gradient of 35%~55% for ultracentirifugation. The virus band formed between 45% and 55% layer was collected after ultracentrifugation with Beckman SW40.1 rotor for 1 hr at 30,000rpm. B. SBV particles purified by ultracentrifugation was stained with 0.5% uranyl acetate and observed by transmission electron microscope. The sizebar indicates 100nm.
 표
Standard curve of the SBV RdRp specific primer sets. A plasmid containing 0.8kb fragment of SBV RdRp was used for dilution series of PCR template. 1x108 plasmid molecules were diluted over a 8-log scale down to 10 molecules. The PCR efficiency, R2, and slope were 88.5%, 0.99, and –3.6, respectively.
표
Standard curve of the SBV RdRp specific primer sets. A plasmid containing 0.8kb fragment of SBV RdRp was used for dilution series of PCR template. 1x108 plasmid molecules were diluted over a 8-log scale down to 10 molecules. The PCR efficiency, R2, and slope were 88.5%, 0.99, and –3.6, respectively.
 표
The SBV concentration-dependent gene expression patterns of invertebrate immune related genes. The X-axis represents the concentration of target genes in each sample by 2(Ct value of target gene – Ct value of ctin gene). Virus; A. cerana carrying h high amount of SBV, Int; A. cerana carrying intermediate amount of SBV, Ctrl; negative control. Based on qPCR study, the Virus carried four times more SBV than Int does.
표
The SBV concentration-dependent gene expression patterns of invertebrate immune related genes. The X-axis represents the concentration of target genes in each sample by 2(Ct value of target gene – Ct value of ctin gene). Virus; A. cerana carrying h high amount of SBV, Int; A. cerana carrying intermediate amount of SBV, Ctrl; negative control. Based on qPCR study, the Virus carried four times more SBV than Int does.
 표
Alignment of the amino acid sequences for AcAcph-1 and known bee venom acid phosphatase Acph-1-like proteins. A vertical arrow indicates the end of the signal peptide. Potential N-glycosylation sites are indicated by crosses. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ710421), Apis mellifera (GenBank accession no. XP_006569977), Bombus impatiens (GenBank accession no. XP_003486582), and Megachile rotundata (GenBank accession no. XP_003706109). The AcAcph-1 sequence was used as the reference for the identity/similarity (Id/Si) values.
표
Alignment of the amino acid sequences for AcAcph-1 and known bee venom acid phosphatase Acph-1-like proteins. A vertical arrow indicates the end of the signal peptide. Potential N-glycosylation sites are indicated by crosses. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ710421), Apis mellifera (GenBank accession no. XP_006569977), Bombus impatiens (GenBank accession no. XP_003486582), and Megachile rotundata (GenBank accession no. XP_003706109). The AcAcph-1 sequence was used as the reference for the identity/similarity (Id/Si) values.
 표
The alignment of the amino acid sequences for AcICK and known ICK peptides. The characteristic cysteine residues are indicated by solid circles. The signal peptide, N-terminal ICK fold, and C-terminal extension are shown. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ530970), A. mellifera (GenBank accession no. XP_006560077), Pogonomyrmex barbatus (GenBank accession no.ADIH01011549), Linepithema humile (GenBank accession no. ADOQ01003051), Solenopsis invicta (GenBank accession no. JQ700385), Acromyrmex echinatior (GenBank accession no.AEVX01007421), Camponotus floridanus (GenBank accession no. AEAB01015096), Atta cephalotes (GenBank accession no. ADTU01022195), and Harpegnathos saltator (GenBank accession no. AEAC01004275). The AcICK sequence was used as the reference for the identity/similarity (Id/Si) values.
표
The alignment of the amino acid sequences for AcICK and known ICK peptides. The characteristic cysteine residues are indicated by solid circles. The signal peptide, N-terminal ICK fold, and C-terminal extension are shown. The sources of the aligned sequences are Apis cerana (this study, GenBank accession no. KJ530970), A. mellifera (GenBank accession no. XP_006560077), Pogonomyrmex barbatus (GenBank accession no.ADIH01011549), Linepithema humile (GenBank accession no. ADOQ01003051), Solenopsis invicta (GenBank accession no. JQ700385), Acromyrmex echinatior (GenBank accession no.AEVX01007421), Camponotus floridanus (GenBank accession no. AEAB01015096), Atta cephalotes (GenBank accession no. ADTU01022195), and Harpegnathos saltator (GenBank accession no. AEAC01004275). The AcICK sequence was used as the reference for the identity/similarity (Id/Si) values.
 표
Transcriptional expression profiles of the A. cerana apoLp-III gene in the fat body of A. cerana worker bees induced by E. coli , B. thuringiensis, or B. bassiana injection. A cerana worker bees injected with PBS were used as injection controls. Total RNA was isolated from the fat body of worker bees at different time points as indicated above the corresponding lanes. A. cerana apoLp-I I I transcripts were analyzed using northern hybridization with radiolabeled A. cerana apoLp-I I I cDNA (upper). The levels of A. cerana apoLp-I I I mRNA are reported as the means of three measurements that were calculated relative to the levels at 0 h before PBS or microbial injection (shown as 100%) (lower). Bars represent the means ± SD.
표
Transcriptional expression profiles of the A. cerana apoLp-III gene in the fat body of A. cerana worker bees induced by E. coli , B. thuringiensis, or B. bassiana injection. A cerana worker bees injected with PBS were used as injection controls. Total RNA was isolated from the fat body of worker bees at different time points as indicated above the corresponding lanes. A. cerana apoLp-I I I transcripts were analyzed using northern hybridization with radiolabeled A. cerana apoLp-I I I cDNA (upper). The levels of A. cerana apoLp-I I I mRNA are reported as the means of three measurements that were calculated relative to the levels at 0 h before PBS or microbial injection (shown as 100%) (lower). Bars represent the means ± SD.
 표
Relationships among the Apis cerana NC2 haplotypes. The tree was constructed using the minimumevolution method that was rooted using two haplotypes of A. koschevnikovi (GenBank accession numbers AB072437 and AB072438; Takahashi et al. 2002) and presented as 50% majority-rule consensus tree. Eight haplotypes were found new in this study (bold), and 48 haplotypes reported previously were included in the analysis. Nodal support was obtained by 1,000 replicates.
표
Relationships among the Apis cerana NC2 haplotypes. The tree was constructed using the minimumevolution method that was rooted using two haplotypes of A. koschevnikovi (GenBank accession numbers AB072437 and AB072438; Takahashi et al. 2002) and presented as 50% majority-rule consensus tree. Eight haplotypes were found new in this study (bold), and 48 haplotypes reported previously were included in the analysis. Nodal support was obtained by 1,000 replicates.
 표
Linear map of Apis florea mitochondrial genome, with the schematic map of triplicated tRNA-Ser(AGN) regionlocated between tRNA-Glu and tRNA-Met and the A+T-rich region. The blank boxes are protein-coding genes that are identical in all hymenopteran family Apidae in arrangement. tRNAs are denoted as one-letter symbols in accordance with the IUPAC-IUB single-letter amino acid codes. The one-letter symbols L, L*, S and S* denote tRNA-Leu(CUN), tRNA-Leu(UUR), tRNA-Ser(AGN), and tRNA-Ser(UCN), respectively. Gene names that are not underlined indicate a clockwise direction of transcription, whereas underlines indicate a counter-clockwise direction. Sequences of triplicated tRNAser(AGN) region are shown with the precedent (RA1 ~ RA3) and following repeats (RB1 ~RB3) plus an extra RA4. SF and LF refer short and long fragments, respectively. LF3 was sequenced by shotgun method, whereas LF1 and LF2 were used as temperate for the amplification and sequencing of the 19 SFs. Schematic map of 1,987-bp long A+T-rich region is composed of two repeat units, named as RC (RC1 ~ RC4) and RD (RD1 ~ RD12) encompassed by three non-repeat sequence (NRS). RD4 ~ RD9 within dotted wavy line were omitted in this map.
표
Linear map of Apis florea mitochondrial genome, with the schematic map of triplicated tRNA-Ser(AGN) regionlocated between tRNA-Glu and tRNA-Met and the A+T-rich region. The blank boxes are protein-coding genes that are identical in all hymenopteran family Apidae in arrangement. tRNAs are denoted as one-letter symbols in accordance with the IUPAC-IUB single-letter amino acid codes. The one-letter symbols L, L*, S and S* denote tRNA-Leu(CUN), tRNA-Leu(UUR), tRNA-Ser(AGN), and tRNA-Ser(UCN), respectively. Gene names that are not underlined indicate a clockwise direction of transcription, whereas underlines indicate a counter-clockwise direction. Sequences of triplicated tRNAser(AGN) region are shown with the precedent (RA1 ~ RA3) and following repeats (RB1 ~RB3) plus an extra RA4. SF and LF refer short and long fragments, respectively. LF3 was sequenced by shotgun method, whereas LF1 and LF2 were used as temperate for the amplification and sequencing of the 19 SFs. Schematic map of 1,987-bp long A+T-rich region is composed of two repeat units, named as RC (RC1 ~ RC4) and RD (RD1 ~ RD12) encompassed by three non-repeat sequence (NRS). RD4 ~ RD9 within dotted wavy line were omitted in this map.
 표
Circular map of the Apis andreniformis mitochondrial genome with the tandemly repeatedtRNA-Leu(CUN) region sequences. Four additional tRNA-Leu(CUN) were found between tRNA-Met and tRNA-Gln, and one additional tRNA-Leu(CUN) was found between tRNA-Gln and tRN-AAla. A nearly identical 68-bp long repeat sequence (repeats 1–6) abutted the 5′-end of each additional tRNA-Leu(CUN) and tRNA-Gln.
표
Circular map of the Apis andreniformis mitochondrial genome with the tandemly repeatedtRNA-Leu(CUN) region sequences. Four additional tRNA-Leu(CUN) were found between tRNA-Met and tRNA-Gln, and one additional tRNA-Leu(CUN) was found between tRNA-Gln and tRN-AAla. A nearly identical 68-bp long repeat sequence (repeats 1–6) abutted the 5′-end of each additional tRNA-Leu(CUN) and tRNA-Gln.
 표
Locations of three major non-coding regions located in the A. cerana mitochondrial genome. Positions ofSer Ile(A) NC1 located between tRNA and tRNA , NC3 located between COIII, and ND3 NC4 located between ND4L and ND6. S, E, M, Q, A, I, G, T and P, respectively, indicate tRNASer, tRNAGlu, tRNAMet, tRNAGln, tRNAAla, tRNAIle, tRNAGly, tRNAThr, and tRNAPro, respectively.
표
Locations of three major non-coding regions located in the A. cerana mitochondrial genome. Positions ofSer Ile(A) NC1 located between tRNA and tRNA , NC3 located between COIII, and ND3 NC4 located between ND4L and ND6. S, E, M, Q, A, I, G, T and P, respectively, indicate tRNASer, tRNAGlu, tRNAMet, tRNAGln, tRNAAla, tRNAIle, tRNAGly, tRNAThr, and tRNAPro, respectively.
| 과제명(ProjectTitle) : | - |
|---|---|
| 연구책임자(Manager) : | - |
| 과제기간(DetailSeriesProject) : | - |
| 총연구비 (DetailSeriesProject) : | - |
| 키워드(keyword) : | - |
| 과제수행기간(LeadAgency) : | - |
| 연구목표(Goal) : | - |
| 연구내용(Abstract) : | - |
| 기대효과(Effect) : | - |
Copyright KISTI. All Rights Reserved.

※ AI-Helper는 부적절한 답변을 할 수 있습니다.