최소 단어 이상 선택하여야 합니다.
최대 10 단어까지만 선택 가능합니다.
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
NTIS 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
DataON 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Edison 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Kafe 바로가기
| 주관연구기관 | 경북대학교 KyungPook National University |
|---|---|
| 연구책임자 | 김정상 |
| 참여연구자 | 김정환 , 박천석 , 임진규 |
| 보고서유형 | 최종보고서 |
| 발행국가 | 대한민국 |
| 언어 | 한국어 |
| 발행년월 | 2010-05 |
| 과제시작연도 | 2010 |
| 주관부처 | 교육과학기술부 Ministry of Education and Science Technology(MEST) |
| 등록번호 | TRKO202100005426 |
| 과제고유번호 | 1345101130 |
| 사업명 | 미래기반기술개발 |
| DB 구축일자 | 2021-07-17 |
| 키워드 | 청국장.된장.메주.바실러스.아스퍼질러스.유전체.단백질체.이소플라본.cheonggukjang.doenjang.meju.Bacillus.Aspergillus.genome.proteome.isoflavones. |
본 세부과제에서는 1단계 연구에서 확립된 유전체 및 단백질체 방법을 이용하여 전통청국장과 메주로부터 우수 균주를 분리동정한 후, 건강기능성과 관련이 깊은 유전자를 클로닝하고 다른 숙주에서의 발현을 시도하였으며, 각 균주들의 콩 발효특성을 연구하여 다음과 같은 주요 결과를 도출하였다.
1. Bacillus 균주에서 polyglutamate(PGA) 생산 4.5 kb pgsBCA 유전자 클로닝 및 유전자 도입에 의한 대장균과 B. subtilis에서 발현
2. 혈전용해효소 유전자 bpr86-1 클로닝 및 B. subtilis
본 세부과제에서는 1단계 연구에서 확립된 유전체 및 단백질체 방법을 이용하여 전통청국장과 메주로부터 우수 균주를 분리동정한 후, 건강기능성과 관련이 깊은 유전자를 클로닝하고 다른 숙주에서의 발현을 시도하였으며, 각 균주들의 콩 발효특성을 연구하여 다음과 같은 주요 결과를 도출하였다.
1. Bacillus 균주에서 polyglutamate(PGA) 생산 4.5 kb pgsBCA 유전자 클로닝 및 유전자 도입에 의한 대장균과 B. subtilis에서 발현
2. 혈전용해효소 유전자 bpr86-1 클로닝 및 B. subtilis에서 생산
3. B. cereus 신속 동정 RAPD-PCR 및 B. subtilis 그룹 종들 구별 RAPD-PCR 방법 개발
4. 전통청국장에서 분리한 Bacillus 균주별 isoflavones대사특성 및 청국장의 건강기능성 규명
5. Bacillus subtilis CH97 와 Bacillus amyloliquefaciens 균주의 secretome 분석완료
6. 전통식품유래 Aspergillus 균주로부터 polyketide 생합성 유전자의 다양성 분석 및 확인
7. 전통식품유래 Aspergillus oryzae 균주로부터 기능성 당으로 그 활용도가 높은 arabinose를 생산하는데 이용할 수 있는 arabinofuranosidase와 기능성 식품소재로 이용이 가능한 ferulate를 생산하는데 사용되는 feruloyl esterase를 cloning하고 효모에서 발현
8. Aspergillus oryzae strains별 beta-glucosidase 활성과 콩 발효과정중 isoflavones 대사 및 항산화 활성에 미치는 영향 평가하여 최적 발효기간 설정
9. Aspergillus oryzae 의 세포질 단백질과 secretome분석하여 부분적인 protein map 완성
10. 단백질분해효소 저항성 콩단백질과 상호작용하는 장점막단백질의 동정을 통한 소화생리학적 새로운 기능에 대한 가능성 제시.
(출처 : 보고서 요약서 3p)
This study was conducted to screen the high quality microorganisms suitable for manufacturing traditional fermented soy foods such as cheonggukjang, soy sauce, and soy paste (doenjang). The genomic and proteomic technologies were utilized for reliable identification of the microorganism. Various str
This study was conducted to screen the high quality microorganisms suitable for manufacturing traditional fermented soy foods such as cheonggukjang, soy sauce, and soy paste (doenjang). The genomic and proteomic technologies were utilized for reliable identification of the microorganism. Various strains in Bacillus and Aspergillus family which were isolated from traditional cheonggukjang and meju, respectively, were tested for their capability to produce soy products with health benefits including antioxidant activity.
Bacilli, major microorganisms found in traditional cheonggukjang, confer important health-related benefits to fermented soybeans. The production of γ-PGA is one example and fibrinolysisis is another. Genes responsible for the γ-PGA production were cloned from B. amyloliquefaciens CH86-1, a strain isolated from traditional cheonggukjang through this study, and the genetic organization was elucidated. A 4.5 kb fragment encompassing pgsBCA was introduced into B. subtilis and E. coli. Production of protein(s) was observed by SDS-PAGE and the presence of transcript was confirmed by slot blot analysis. More detailed studies are required for transforming a PGA-nonproducer into an efficient producer. Fibrinolytic enzymes were studied more in detail and bpr86-1 gene encoding bacillopeptidase F was cloned from B. amyloiliquefaciens CH86-1. These genes will be useful for the construction of strains for fermented soy products with enhanced health benefits.
A rapid and reliable RAPD-PCR method were also developed for the detection of Bacillus cereus, a common contaminant, from fermented soyfoods. All B. cereus strains produced 0.91 kb and 0.5 kb when S-30 was used as a single primer. A RAPD-PCR protocol was also developed to distinguish B. subtilis strains from closely related B. amyloliquefaciens and B. licheniformis strains. Accurate identification of a Bacillus species is helpful for the quality control of fermented foods.
Meanwhile, five Aspergillus oryzae strains were selected from Korean traditional meju manufactured in Sunchang Yeastopia by ITS-RFLP method. The absence of aflatoxin biosynthesis by five selected Aspergillus oryzae strains was confirmed by using RT-PCR. In addition, 8 β-glucosidase genes (ABG 1, 7, 9, 13, 21, 22, 30, and 31) from one of selected Asp oryzae strain were cloned and expressed in Saccharomyces cerevisiae . The β-glucosidase expressed on the yeast could hydrolyze various isoflavones in soybean. Although five strains of Aspergillus oryzae expressed different levels of the enzymes, they did not have significant effect on the profile of isoflavone derivatives during soybean fermentation. However, isoflavone profile of soybean was greatly altered depending upon soybean variety and fermentation period, with the highest concentration of free isoflavones at 4-5th day after fermentation in high isoflavone variety (Aga 3). The antioxidant activity of fermented soy was proportional to the free isoflavone content.
Two biotechnologically important enzymes, arabinofuranosidase and feruloyl esterase, were cloned and expressed in Pichia pastoris and their properties were characterized. This research also undertook the proteome analysis of Bacillus and Aspergillus strains isolated from the traditionally prepared cheonggukjang and meju, proteome analysis of the intestinal mucosa from the rats fed with cheonggukjang, and observed the interaction of the pepsin-resistant soy proteins with the intestinal mucosa membrane proteins as well. Proteomes of Bacillus subtilis and B. amyloliquefaciens were separated on 2-D gels and identified by peptide mass fingerprinting. Due to the variations in genomic sequences, the identified proteins were overlapped less than 10%. The phenomenon is also true between same species. The proteome profiles obtained from the bacillus strains could be used to classify the microorganisms isolated throughout this project. In case of Aspergillus, 2-D proteome profiles agree each other very well within the same species (over 90%), such as between Aspergillus oryzae strains. However, proteome profiles between A. oryzae matched only by 50% and A. awamori only overlapped by less than 20%. This also suggests that the Aspergillus strains identified by genomic sequence data should be complemented by proteomic data for accurate identification and classification.
Proteomic analysis of the mucosa of the small intestine from the rats fed with cheonggukjang and soy showed differential expression of cytoskeletal proteins, particularly, tropomyosin, which cause the increase of the length of microvilli of the small intestine. This phenomenon could be caused by the interaction of the pepsin-resistant soy proteins with intestinal surface. The pepsin-resistant soy proteins were found to interact with a 32 kDa protein on the mucosa membrane. The interaction of soy proteins with small intestinal mucosa need to be further investigated.
In conclusion, we made major breakthrough in understanding microorganisms involved in manufacturing Korean traditional fermented soy foods. The information generated from this study should be exploited for improving manufacturing process of traditional fermented soy products.
(출처 : SUMMARY 9p)
 표
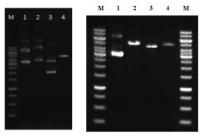
Restriction cut of vectors and pgsBCA genes. (left) 1. pHY300PLK, 2. pET-26b(+), 3, pBlueScript, 4. pgs genes amplified (5.3 kb) (right) 1. pHY300PLK, 2. gel extracted pgs gens after amplification (5.3 kb), 3. pHY300PLK, SalI and BamHI cut, 4. pgs genes, SalI and BamHI cut M, size maker (Fermentas, Burlington, Canada)
표
Restriction cut of vectors and pgsBCA genes. (left) 1. pHY300PLK, 2. pET-26b(+), 3, pBlueScript, 4. pgs genes amplified (5.3 kb) (right) 1. pHY300PLK, 2. gel extracted pgs gens after amplification (5.3 kb), 3. pHY300PLK, SalI and BamHI cut, 4. pgs genes, SalI and BamHI cut M, size maker (Fermentas, Burlington, Canada)
![SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers](https://nrms.kisti.re.kr/bitextimages/TRKO202100005426/TRKO202100005426_50_image_2.png) 표
SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers
표
SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers
![Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control](https://nrms.kisti.re.kr/bitextimages/TRKO202100005426/TRKO202100005426_55_image_1.png) 표
Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control
표
Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control
![SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control](https://nrms.kisti.re.kr/bitextimages/TRKO202100005426/TRKO202100005426_55_image_2.png) 표
SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control
표
SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control
 표
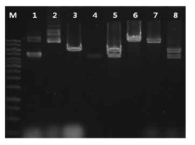
Restriction digestion of pGHk prepared from B. subtilis TF. M, 1 kb size maker (Fermentas); 1, pHY3-5 plasmid DNA from DH5α; 2, pGHk plasmid DNA from B. subtilis WB600; 3, #1 digested with BamHI (6.6 kb); 4, bpr86-1 gene (insert, 4.99 kb); 5, pGHk plasmid DNA (from DH5α) digested with XbaI and BamH (6.6 kb, 4.99 kb); 6, pGHk plasmid DNA (from DH5α) digested with BamHI (11.6 kb); 7, #2 digested with BamHI (11.6 kb); 8, #2 digested with XbaI and BamH (6.6 kb, 4.99 kb)
표
Restriction digestion of pGHk prepared from B. subtilis TF. M, 1 kb size maker (Fermentas); 1, pHY3-5 plasmid DNA from DH5α; 2, pGHk plasmid DNA from B. subtilis WB600; 3, #1 digested with BamHI (6.6 kb); 4, bpr86-1 gene (insert, 4.99 kb); 5, pGHk plasmid DNA (from DH5α) digested with XbaI and BamH (6.6 kb, 4.99 kb); 6, pGHk plasmid DNA (from DH5α) digested with BamHI (11.6 kb); 7, #2 digested with BamHI (11.6 kb); 8, #2 digested with XbaI and BamH (6.6 kb, 4.99 kb)
 표
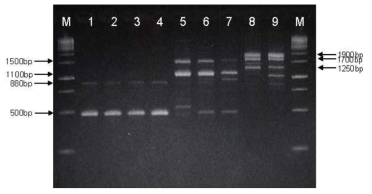
RAPD-PCR patterns of Bacillus strains isolated from Chunggukjang and reference strains. Lane: 1, B. subtilis168; 2, CH3-5; 3, CH3-25; 4, CH97; 5, B. amyloliquefaciens ATCC 23845; 6, CH51; 7, CH86-1; 8, B. licheniformis ATCC 27811; 9, CH3-17; M, 1 kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used for the electrophoresis
표
RAPD-PCR patterns of Bacillus strains isolated from Chunggukjang and reference strains. Lane: 1, B. subtilis168; 2, CH3-5; 3, CH3-25; 4, CH97; 5, B. amyloliquefaciens ATCC 23845; 6, CH51; 7, CH86-1; 8, B. licheniformis ATCC 27811; 9, CH3-17; M, 1 kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used for the electrophoresis
 표
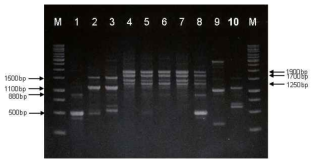
RAPD-PCR patterns of Bacillus reference strains. Lane: 1, B. subtilis 168; 2, B. amyloliquefaciensATCC 23845; 3, B. amyloliquefaciens ATCC 23350; 4, B. licheniformis ATCC 4527; 5, B. licheniformis ATCC 21416; 6, B. licheniformis ATCC 27811; 7, B. licheniformis ATCC39326; 8, B. licheniformisATCC 14580; 9, B. thuringiensis ATCC 33679; 10, B. circulans ATCC 4513; M, 1kb size ladder (MBI, Glen Burnie, USA).2% agarose gel was used
표
RAPD-PCR patterns of Bacillus reference strains. Lane: 1, B. subtilis 168; 2, B. amyloliquefaciensATCC 23845; 3, B. amyloliquefaciens ATCC 23350; 4, B. licheniformis ATCC 4527; 5, B. licheniformis ATCC 21416; 6, B. licheniformis ATCC 27811; 7, B. licheniformis ATCC39326; 8, B. licheniformisATCC 14580; 9, B. thuringiensis ATCC 33679; 10, B. circulans ATCC 4513; M, 1kb size ladder (MBI, Glen Burnie, USA).2% agarose gel was used
 표
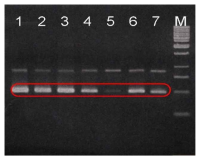
RAPD-PCR profiles of seven B. subtilis reference strains. All seven B. subtilis reference strains showed the same pattern and 500 bp fragments were circled. Lane: 1, B. subtilis 168; 2, B. subtilis ATCC 33608; 3, B. subtilisATCC 6051A; 4, B. subtilis ATCC 9372; 5, B. subtilis ATCC 39088; 6, B. subtilisATCC 33234; 7, B. subtilis ATCC 21777; M, 1kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used
표
RAPD-PCR profiles of seven B. subtilis reference strains. All seven B. subtilis reference strains showed the same pattern and 500 bp fragments were circled. Lane: 1, B. subtilis 168; 2, B. subtilis ATCC 33608; 3, B. subtilisATCC 6051A; 4, B. subtilis ATCC 9372; 5, B. subtilis ATCC 39088; 6, B. subtilisATCC 33234; 7, B. subtilis ATCC 21777; M, 1kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used
 표
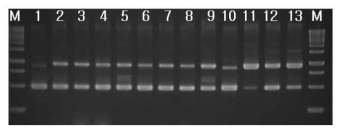
RAPD-PCR of 3 strains isolated from cheonggukjang and standard strains. 1, B. subtilis 168; 2, B. cereus ATCC9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, 부산대 1; 12. 부산대 2; 13, 부산대 3; M, 1kb size ladder (MBI). 2% agarose gel이 사용되었다
표
RAPD-PCR of 3 strains isolated from cheonggukjang and standard strains. 1, B. subtilis 168; 2, B. cereus ATCC9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, 부산대 1; 12. 부산대 2; 13, 부산대 3; M, 1kb size ladder (MBI). 2% agarose gel이 사용되었다
 표
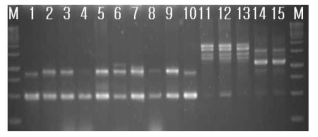
RAPD-PCR results of standard Bacillus strains. 1, B. subtilis 168; 2, B. cereus ATCC 9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, B. licheniformis ATCC4527; 12. B. licheniformis ATCC21416; 13, B. licheniformis ATCC27811; 14, B. amyloliquefaciens ATCC23350; 15, B. amyloliquefaciens ATCC23845; M, 1kb size ladder (MBI). 2% agarose gel was used
표
RAPD-PCR results of standard Bacillus strains. 1, B. subtilis 168; 2, B. cereus ATCC 9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, B. licheniformis ATCC4527; 12. B. licheniformis ATCC21416; 13, B. licheniformis ATCC27811; 14, B. amyloliquefaciens ATCC23350; 15, B. amyloliquefaciens ATCC23845; M, 1kb size ladder (MBI). 2% agarose gel was used
 표
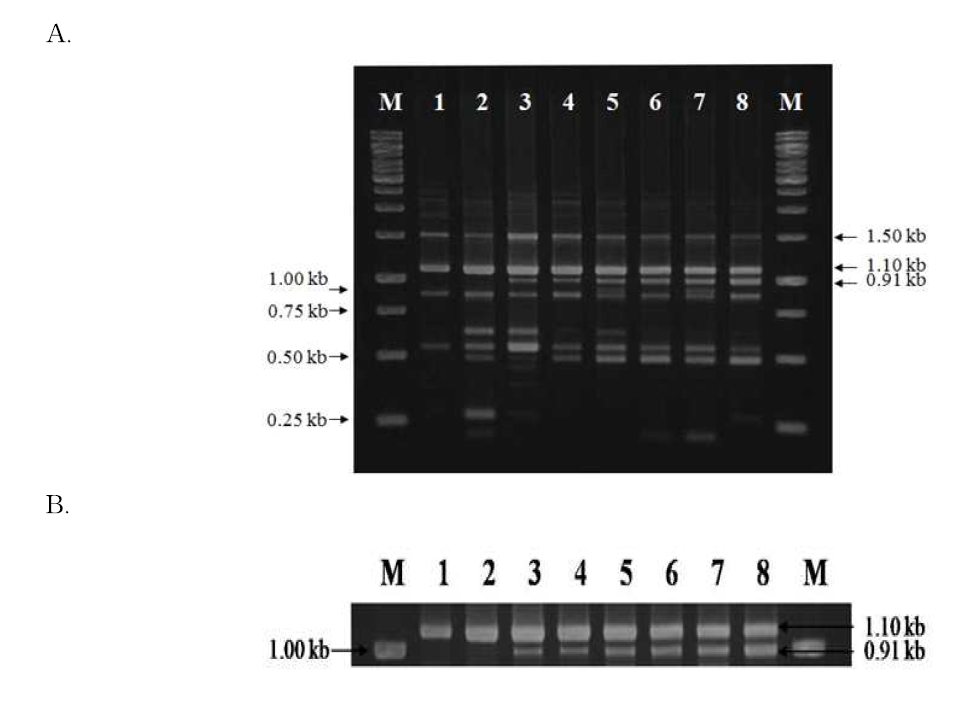
RAPD-PCR patterns of cheonggukjang spiked with B. cereus. A. 1, control (non-spiked cheonggukjang) 2-8, cheonggukjang samples spiked with B. cereus cells (cells/g cheongukjang): 2, 100 ; 3, 101 4, 102 5, 103 6, 104 ;7, 105 8, 106 , M, 1 kb size ladder (MBI Fermentas). B. Enlarged picture showing the 0.91 kb fragment
표
RAPD-PCR patterns of cheonggukjang spiked with B. cereus. A. 1, control (non-spiked cheonggukjang) 2-8, cheonggukjang samples spiked with B. cereus cells (cells/g cheongukjang): 2, 100 ; 3, 101 4, 102 5, 103 6, 104 ;7, 105 8, 106 , M, 1 kb size ladder (MBI Fermentas). B. Enlarged picture showing the 0.91 kb fragment
 표
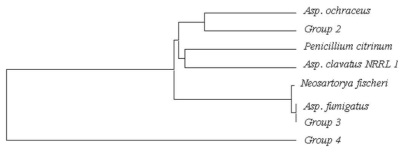
Neighbor-joining phylogenetic tree of HR KS domain PKS (Group 2, Asp. oryzae; Group 3, Asp. fumigatus; Group 4, Arthrinium sp.) with Asp. ochraceus polyketide synthase 1, Asp. fumigatus LovB-like polyketide synthase, Asp. clavatus NRRL1 equisetin synthetase, Neosartorya fischeri Polyketide synthase, and Penicillium citrinum Polyketide synthase
표
Neighbor-joining phylogenetic tree of HR KS domain PKS (Group 2, Asp. oryzae; Group 3, Asp. fumigatus; Group 4, Arthrinium sp.) with Asp. ochraceus polyketide synthase 1, Asp. fumigatus LovB-like polyketide synthase, Asp. clavatus NRRL1 equisetin synthetase, Neosartorya fischeri Polyketide synthase, and Penicillium citrinum Polyketide synthase
 표
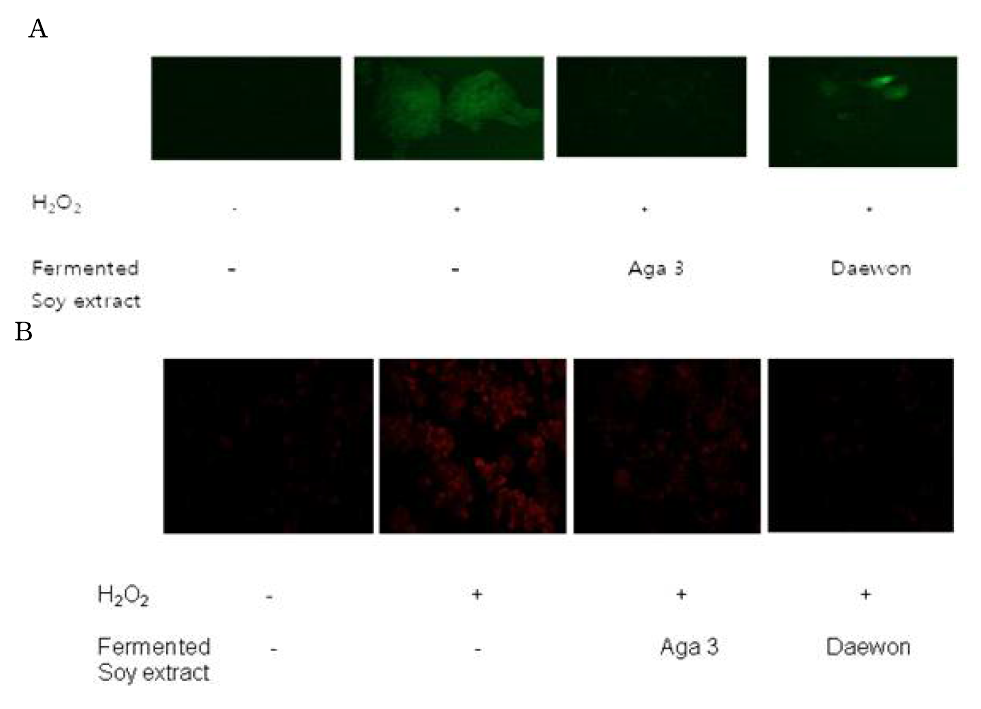
Detection of intracellular reactive oxygen species by DCFDA (A) and DHE (B) in hepa1c1c7 cells. Hepa1c1c7 cells were exposed to hydrogen peroxide in the absence and presence of fermented soy extract from Aga 3 or Daewon variety of soybean, followed by observation under fluorescent microscope after the addition of DCFDA or DHE, fluorescent probes
표
Detection of intracellular reactive oxygen species by DCFDA (A) and DHE (B) in hepa1c1c7 cells. Hepa1c1c7 cells were exposed to hydrogen peroxide in the absence and presence of fermented soy extract from Aga 3 or Daewon variety of soybean, followed by observation under fluorescent microscope after the addition of DCFDA or DHE, fluorescent probes
 표
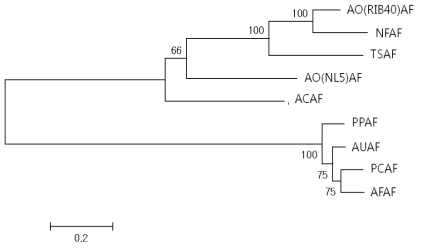
Phylogenetic tree of putative arabinofuranosidase genes, AO(RIB40)AF, AFase from Aspergillus oryzae RIB40; AFase from Neosartorya fischeri NRRL 181-5; TSAF, AFase from Talaromyces stipitatus ATCC 10500; AO(NL5)AF AFase from Aspergillus oryzae FMB-2a; ACAF, AFase from Aspergillus clavatus NRRL 1-3; PPAF. AFase from Penicillium purpurogenum; AUAF, AFase from Aspergillus fumigatus Af293; PCAF, AFase from Penicillium chrysogenum-6; AFAF, AFase from Aspergillus flavus NRRL3357
표
Phylogenetic tree of putative arabinofuranosidase genes, AO(RIB40)AF, AFase from Aspergillus oryzae RIB40; AFase from Neosartorya fischeri NRRL 181-5; TSAF, AFase from Talaromyces stipitatus ATCC 10500; AO(NL5)AF AFase from Aspergillus oryzae FMB-2a; ACAF, AFase from Aspergillus clavatus NRRL 1-3; PPAF. AFase from Penicillium purpurogenum; AUAF, AFase from Aspergillus fumigatus Af293; PCAF, AFase from Penicillium chrysogenum-6; AFAF, AFase from Aspergillus flavus NRRL3357
 표
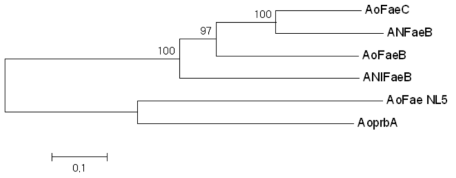
Phylogenetic tree of putative feruloyl esterase genes, AoFaeC, Aspergillus oryzae feruloyl esterase C (XP_001819091); ANFaeB, Aspergillus nidulans feruloyl esterase (XP_659376); AoFaeB, Aspergillus oryzae feruloyl esterase B (XP_001818628); ANIFaeB, Aspergillus niger feruloyl esterase(CAC83933); AoFaeNL5, Aspergillus oryzae FMB-2a feruloyl esterase; AoprbA, Aspergillus oryzae feruloyl esterase B (XP_001821143)
표
Phylogenetic tree of putative feruloyl esterase genes, AoFaeC, Aspergillus oryzae feruloyl esterase C (XP_001819091); ANFaeB, Aspergillus nidulans feruloyl esterase (XP_659376); AoFaeB, Aspergillus oryzae feruloyl esterase B (XP_001818628); ANIFaeB, Aspergillus niger feruloyl esterase(CAC83933); AoFaeNL5, Aspergillus oryzae FMB-2a feruloyl esterase; AoprbA, Aspergillus oryzae feruloyl esterase B (XP_001821143)
 표
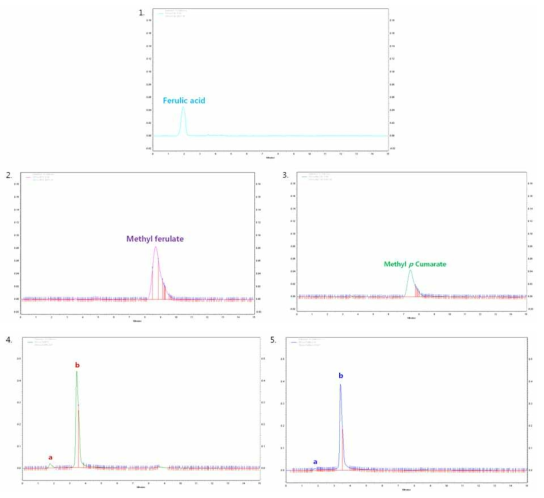
HPLC analysis of FEase reaction products. Lane 1, Ferulic acid control; Lane 2, Methyl ferulate control; Lane 3, methyl-p-coumarate control; Lane 4, reaction products with methyl ferulate as substrate, a, ferulic acid; b, p-coumaric acid; Lane 5, reaction products with methyl-p-coumarate as substrate
표
HPLC analysis of FEase reaction products. Lane 1, Ferulic acid control; Lane 2, Methyl ferulate control; Lane 3, methyl-p-coumarate control; Lane 4, reaction products with methyl ferulate as substrate, a, ferulic acid; b, p-coumaric acid; Lane 5, reaction products with methyl-p-coumarate as substrate
 표
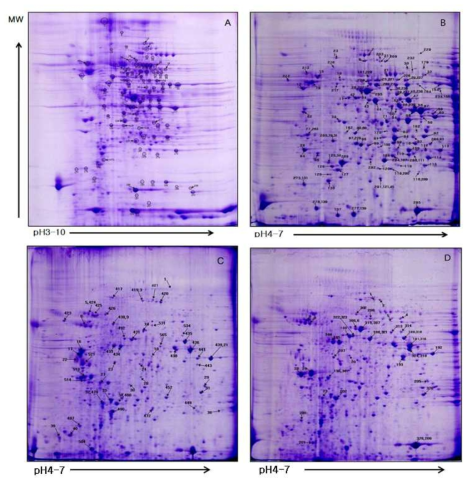
2-D gel containing intestinal proteins from a pool of designated group of small intestinal mucosa samples stained with coomassie blue. All IEF was perfomed on an 17-cm length of IPG strip having a pH range of 3-10 (linear pH gradient). The second separation was done using a 12% SDS-PAGE gel; A. control, B. cheongkukjang, C. control, D. soy E. genistein, F. control
표
2-D gel containing intestinal proteins from a pool of designated group of small intestinal mucosa samples stained with coomassie blue. All IEF was perfomed on an 17-cm length of IPG strip having a pH range of 3-10 (linear pH gradient). The second separation was done using a 12% SDS-PAGE gel; A. control, B. cheongkukjang, C. control, D. soy E. genistein, F. control
 표
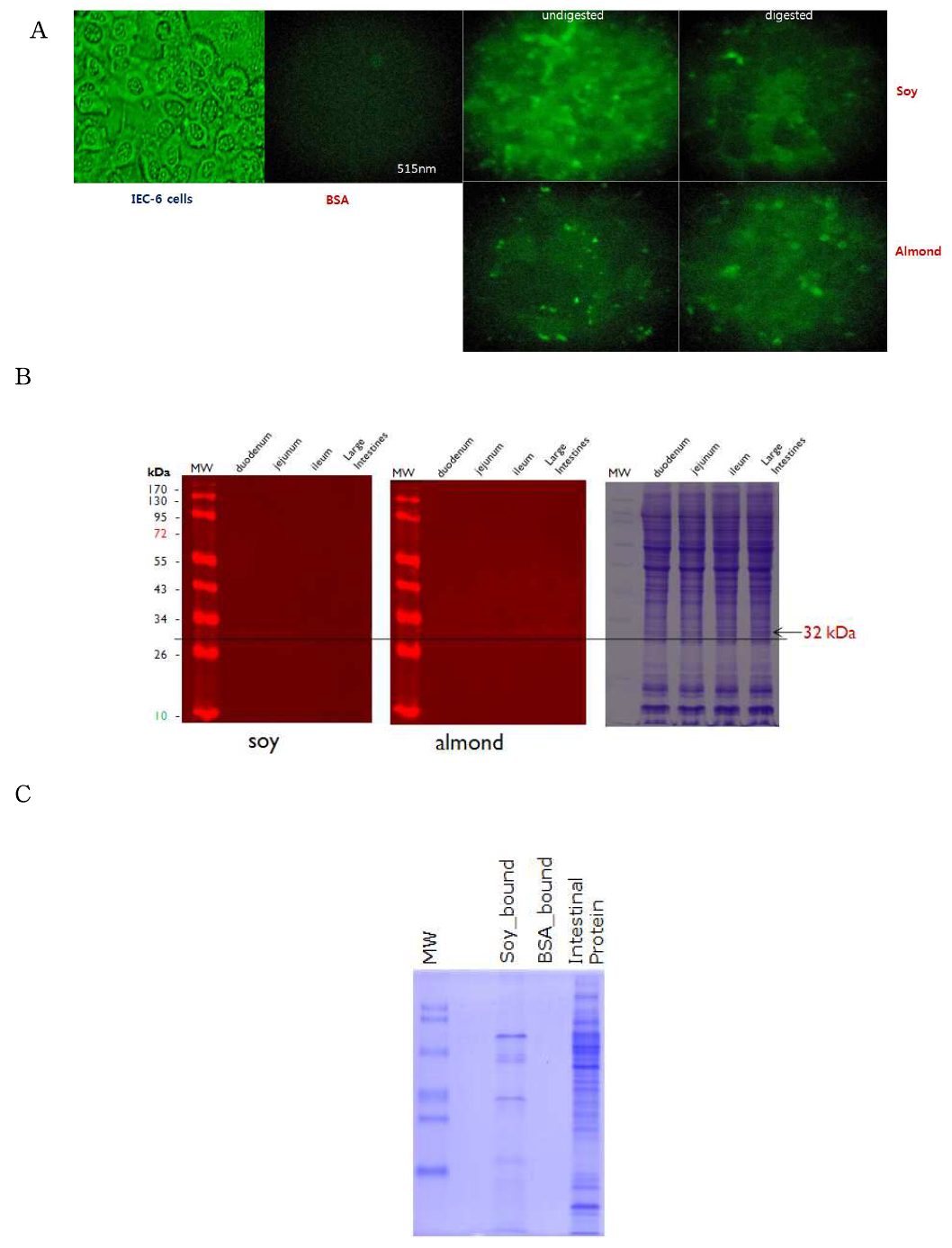
Ligand binding on FITC-labed protein of IEC cells. 콩이나 아몬드 단백질들은 BSA 와 비교할 때 현저하게 높은 결합을 보였으며 digested 또는 undigested 단백질들이 모두 결합됨을 알 수 있다 (A). 콩과 아몬드 단백질이 소장점막 단백질 중 32 kDa 단백질에 특이적으로 결합하였다 (B). Soy protein이 결합된 affinity column을 만들어서 장점막단백질과의 결합을 이루고 콩단백질과 결합하는 장점막단백질을 분리하여 1-D SDS-PAGE로 단백질을 분리하였다(C)
표
Ligand binding on FITC-labed protein of IEC cells. 콩이나 아몬드 단백질들은 BSA 와 비교할 때 현저하게 높은 결합을 보였으며 digested 또는 undigested 단백질들이 모두 결합됨을 알 수 있다 (A). 콩과 아몬드 단백질이 소장점막 단백질 중 32 kDa 단백질에 특이적으로 결합하였다 (B). Soy protein이 결합된 affinity column을 만들어서 장점막단백질과의 결합을 이루고 콩단백질과 결합하는 장점막단백질을 분리하여 1-D SDS-PAGE로 단백질을 분리하였다(C)
 표
Restriction cut of vectors and pgsBCA genes. (left) 1. pHY300PLK, 2. pET-26b(+), 3, pBlueScript, 4. pgs genes amplified (5.3 kb) (right) 1. pHY300PLK, 2. gel extracted pgs gens after amplification (5.3 kb), 3. pHY300PLK, SalI and BamHI cut, 4. pgs genes, SalI and BamHI cut M, size maker (Fermentas, Burlington, Canada)
표
Restriction cut of vectors and pgsBCA genes. (left) 1. pHY300PLK, 2. pET-26b(+), 3, pBlueScript, 4. pgs genes amplified (5.3 kb) (right) 1. pHY300PLK, 2. gel extracted pgs gens after amplification (5.3 kb), 3. pHY300PLK, SalI and BamHI cut, 4. pgs genes, SalI and BamHI cut M, size maker (Fermentas, Burlington, Canada)
![SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers](https://nrms.kisti.re.kr/bitextimages/TRKO202100005426/TRKO202100005426_50_image_2.png) 표
SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers
표
SDS-PAGE for protein samples from E. coli BL21(DE3)[pGAk100]. Lane 1-4, soluble proteins and lane 5-8, insoluble proteins. BL21 [pET-26b(+)] at 0.1 mM IPTG(1 and 5); BL21 [pGAk100] at 0 mM IPTG (2 and 6); BL21[pGAk100] at 0.1 mM IPTG (3 and 7); BL21[pGAk100] at 1 mM IPTG (4 and 8). M, size markers
![Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control](https://nrms.kisti.re.kr/bitextimages/TRKO202100005426/TRKO202100005426_55_image_1.png) 표
Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control
표
Fibrin plate assay showing fibrinolytic activity of B. subtilis WB600[pHYbpr86-1] during growth at 37 ℃. -○-, B. subtilis WB600[pHY300PLK]; -● -, B. subtilis WB600[pHYbpr86-1]. Culture supernatant at each time point was obtained, filtered, and then spotted on a fibrin plate. Plasmin at different concentration was used as a positive control
![SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control](https://nrms.kisti.re.kr/bitextimages/TRKO202100005426/TRKO202100005426_55_image_2.png) 표
SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control
표
SDS-PAGE (left) and fibrin zymogram (right) of culture supernatant. M, protein size markers(DokDo-MARKTM, Elpisbio, Taejeon, Korea); B. amyloliquefaciens CH86-1 at 48 h (lane 1), at 96 h (2), at 120 h (3); B. subtilis WB600[pHYbpr86-1] at 48 h (4), at 96 h (5), at 120 h (6); 7, B. subtilis WB600[pHY300PLK] at 48 h (7), at 96 h (8), at 120 h (9). 15 μg was applied and 10% acrylamide gel was used. Bands marked in the left are absent in control
 표
Restriction digestion of pGHk prepared from B. subtilis TF. M, 1 kb size maker (Fermentas); 1, pHY3-5 plasmid DNA from DH5α; 2, pGHk plasmid DNA from B. subtilis WB600; 3, #1 digested with BamHI (6.6 kb); 4, bpr86-1 gene (insert, 4.99 kb); 5, pGHk plasmid DNA (from DH5α) digested with XbaI and BamH (6.6 kb, 4.99 kb); 6, pGHk plasmid DNA (from DH5α) digested with BamHI (11.6 kb); 7, #2 digested with BamHI (11.6 kb); 8, #2 digested with XbaI and BamH (6.6 kb, 4.99 kb)
표
Restriction digestion of pGHk prepared from B. subtilis TF. M, 1 kb size maker (Fermentas); 1, pHY3-5 plasmid DNA from DH5α; 2, pGHk plasmid DNA from B. subtilis WB600; 3, #1 digested with BamHI (6.6 kb); 4, bpr86-1 gene (insert, 4.99 kb); 5, pGHk plasmid DNA (from DH5α) digested with XbaI and BamH (6.6 kb, 4.99 kb); 6, pGHk plasmid DNA (from DH5α) digested with BamHI (11.6 kb); 7, #2 digested with BamHI (11.6 kb); 8, #2 digested with XbaI and BamH (6.6 kb, 4.99 kb)
 표
RAPD-PCR patterns of Bacillus strains isolated from Chunggukjang and reference strains. Lane: 1, B. subtilis168; 2, CH3-5; 3, CH3-25; 4, CH97; 5, B. amyloliquefaciens ATCC 23845; 6, CH51; 7, CH86-1; 8, B. licheniformis ATCC 27811; 9, CH3-17; M, 1 kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used for the electrophoresis
표
RAPD-PCR patterns of Bacillus strains isolated from Chunggukjang and reference strains. Lane: 1, B. subtilis168; 2, CH3-5; 3, CH3-25; 4, CH97; 5, B. amyloliquefaciens ATCC 23845; 6, CH51; 7, CH86-1; 8, B. licheniformis ATCC 27811; 9, CH3-17; M, 1 kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used for the electrophoresis
 표
RAPD-PCR patterns of Bacillus reference strains. Lane: 1, B. subtilis 168; 2, B. amyloliquefaciensATCC 23845; 3, B. amyloliquefaciens ATCC 23350; 4, B. licheniformis ATCC 4527; 5, B. licheniformis ATCC 21416; 6, B. licheniformis ATCC 27811; 7, B. licheniformis ATCC39326; 8, B. licheniformisATCC 14580; 9, B. thuringiensis ATCC 33679; 10, B. circulans ATCC 4513; M, 1kb size ladder (MBI, Glen Burnie, USA).2% agarose gel was used
표
RAPD-PCR patterns of Bacillus reference strains. Lane: 1, B. subtilis 168; 2, B. amyloliquefaciensATCC 23845; 3, B. amyloliquefaciens ATCC 23350; 4, B. licheniformis ATCC 4527; 5, B. licheniformis ATCC 21416; 6, B. licheniformis ATCC 27811; 7, B. licheniformis ATCC39326; 8, B. licheniformisATCC 14580; 9, B. thuringiensis ATCC 33679; 10, B. circulans ATCC 4513; M, 1kb size ladder (MBI, Glen Burnie, USA).2% agarose gel was used
 표
RAPD-PCR profiles of seven B. subtilis reference strains. All seven B. subtilis reference strains showed the same pattern and 500 bp fragments were circled. Lane: 1, B. subtilis 168; 2, B. subtilis ATCC 33608; 3, B. subtilisATCC 6051A; 4, B. subtilis ATCC 9372; 5, B. subtilis ATCC 39088; 6, B. subtilisATCC 33234; 7, B. subtilis ATCC 21777; M, 1kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used
표
RAPD-PCR profiles of seven B. subtilis reference strains. All seven B. subtilis reference strains showed the same pattern and 500 bp fragments were circled. Lane: 1, B. subtilis 168; 2, B. subtilis ATCC 33608; 3, B. subtilisATCC 6051A; 4, B. subtilis ATCC 9372; 5, B. subtilis ATCC 39088; 6, B. subtilisATCC 33234; 7, B. subtilis ATCC 21777; M, 1kb size ladder (MBI, Glen Burnie, USA). 2% agarose gel was used
 표
RAPD-PCR of 3 strains isolated from cheonggukjang and standard strains. 1, B. subtilis 168; 2, B. cereus ATCC9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, 부산대 1; 12. 부산대 2; 13, 부산대 3; M, 1kb size ladder (MBI). 2% agarose gel이 사용되었다
표
RAPD-PCR of 3 strains isolated from cheonggukjang and standard strains. 1, B. subtilis 168; 2, B. cereus ATCC9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, 부산대 1; 12. 부산대 2; 13, 부산대 3; M, 1kb size ladder (MBI). 2% agarose gel이 사용되었다
 표
RAPD-PCR results of standard Bacillus strains. 1, B. subtilis 168; 2, B. cereus ATCC 9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, B. licheniformis ATCC4527; 12. B. licheniformis ATCC21416; 13, B. licheniformis ATCC27811; 14, B. amyloliquefaciens ATCC23350; 15, B. amyloliquefaciens ATCC23845; M, 1kb size ladder (MBI). 2% agarose gel was used
표
RAPD-PCR results of standard Bacillus strains. 1, B. subtilis 168; 2, B. cereus ATCC 9634; 3, B. cereus ATCC21768; 4, B. cereus ATCC14579; 5, B. cereus ATCC31429; 6, B. cereus ATCC27348; 7, B. cereus ATCC21366; 8, B. cereus ATCC21772; 9, B. cereus ATCC12480; 10, B. subtilis 168; 11, B. licheniformis ATCC4527; 12. B. licheniformis ATCC21416; 13, B. licheniformis ATCC27811; 14, B. amyloliquefaciens ATCC23350; 15, B. amyloliquefaciens ATCC23845; M, 1kb size ladder (MBI). 2% agarose gel was used
 표
RAPD-PCR patterns of cheonggukjang spiked with B. cereus. A. 1, control (non-spiked cheonggukjang) 2-8, cheonggukjang samples spiked with B. cereus cells (cells/g cheongukjang): 2, 100 ; 3, 101 4, 102 5, 103 6, 104 ;7, 105 8, 106 , M, 1 kb size ladder (MBI Fermentas). B. Enlarged picture showing the 0.91 kb fragment
표
RAPD-PCR patterns of cheonggukjang spiked with B. cereus. A. 1, control (non-spiked cheonggukjang) 2-8, cheonggukjang samples spiked with B. cereus cells (cells/g cheongukjang): 2, 100 ; 3, 101 4, 102 5, 103 6, 104 ;7, 105 8, 106 , M, 1 kb size ladder (MBI Fermentas). B. Enlarged picture showing the 0.91 kb fragment
 표
Neighbor-joining phylogenetic tree of HR KS domain PKS (Group 2, Asp. oryzae; Group 3, Asp. fumigatus; Group 4, Arthrinium sp.) with Asp. ochraceus polyketide synthase 1, Asp. fumigatus LovB-like polyketide synthase, Asp. clavatus NRRL1 equisetin synthetase, Neosartorya fischeri Polyketide synthase, and Penicillium citrinum Polyketide synthase
표
Neighbor-joining phylogenetic tree of HR KS domain PKS (Group 2, Asp. oryzae; Group 3, Asp. fumigatus; Group 4, Arthrinium sp.) with Asp. ochraceus polyketide synthase 1, Asp. fumigatus LovB-like polyketide synthase, Asp. clavatus NRRL1 equisetin synthetase, Neosartorya fischeri Polyketide synthase, and Penicillium citrinum Polyketide synthase
 표
Detection of intracellular reactive oxygen species by DCFDA (A) and DHE (B) in hepa1c1c7 cells. Hepa1c1c7 cells were exposed to hydrogen peroxide in the absence and presence of fermented soy extract from Aga 3 or Daewon variety of soybean, followed by observation under fluorescent microscope after the addition of DCFDA or DHE, fluorescent probes
표
Detection of intracellular reactive oxygen species by DCFDA (A) and DHE (B) in hepa1c1c7 cells. Hepa1c1c7 cells were exposed to hydrogen peroxide in the absence and presence of fermented soy extract from Aga 3 or Daewon variety of soybean, followed by observation under fluorescent microscope after the addition of DCFDA or DHE, fluorescent probes
 표
Phylogenetic tree of putative arabinofuranosidase genes, AO(RIB40)AF, AFase from Aspergillus oryzae RIB40; AFase from Neosartorya fischeri NRRL 181-5; TSAF, AFase from Talaromyces stipitatus ATCC 10500; AO(NL5)AF AFase from Aspergillus oryzae FMB-2a; ACAF, AFase from Aspergillus clavatus NRRL 1-3; PPAF. AFase from Penicillium purpurogenum; AUAF, AFase from Aspergillus fumigatus Af293; PCAF, AFase from Penicillium chrysogenum-6; AFAF, AFase from Aspergillus flavus NRRL3357
표
Phylogenetic tree of putative arabinofuranosidase genes, AO(RIB40)AF, AFase from Aspergillus oryzae RIB40; AFase from Neosartorya fischeri NRRL 181-5; TSAF, AFase from Talaromyces stipitatus ATCC 10500; AO(NL5)AF AFase from Aspergillus oryzae FMB-2a; ACAF, AFase from Aspergillus clavatus NRRL 1-3; PPAF. AFase from Penicillium purpurogenum; AUAF, AFase from Aspergillus fumigatus Af293; PCAF, AFase from Penicillium chrysogenum-6; AFAF, AFase from Aspergillus flavus NRRL3357
 표
Phylogenetic tree of putative feruloyl esterase genes, AoFaeC, Aspergillus oryzae feruloyl esterase C (XP_001819091); ANFaeB, Aspergillus nidulans feruloyl esterase (XP_659376); AoFaeB, Aspergillus oryzae feruloyl esterase B (XP_001818628); ANIFaeB, Aspergillus niger feruloyl esterase(CAC83933); AoFaeNL5, Aspergillus oryzae FMB-2a feruloyl esterase; AoprbA, Aspergillus oryzae feruloyl esterase B (XP_001821143)
표
Phylogenetic tree of putative feruloyl esterase genes, AoFaeC, Aspergillus oryzae feruloyl esterase C (XP_001819091); ANFaeB, Aspergillus nidulans feruloyl esterase (XP_659376); AoFaeB, Aspergillus oryzae feruloyl esterase B (XP_001818628); ANIFaeB, Aspergillus niger feruloyl esterase(CAC83933); AoFaeNL5, Aspergillus oryzae FMB-2a feruloyl esterase; AoprbA, Aspergillus oryzae feruloyl esterase B (XP_001821143)
 표
HPLC analysis of FEase reaction products. Lane 1, Ferulic acid control; Lane 2, Methyl ferulate control; Lane 3, methyl-p-coumarate control; Lane 4, reaction products with methyl ferulate as substrate, a, ferulic acid; b, p-coumaric acid; Lane 5, reaction products with methyl-p-coumarate as substrate
표
HPLC analysis of FEase reaction products. Lane 1, Ferulic acid control; Lane 2, Methyl ferulate control; Lane 3, methyl-p-coumarate control; Lane 4, reaction products with methyl ferulate as substrate, a, ferulic acid; b, p-coumaric acid; Lane 5, reaction products with methyl-p-coumarate as substrate
 표
2-D gel containing intestinal proteins from a pool of designated group of small intestinal mucosa samples stained with coomassie blue. All IEF was perfomed on an 17-cm length of IPG strip having a pH range of 3-10 (linear pH gradient). The second separation was done using a 12% SDS-PAGE gel; A. control, B. cheongkukjang, C. control, D. soy E. genistein, F. control
표
2-D gel containing intestinal proteins from a pool of designated group of small intestinal mucosa samples stained with coomassie blue. All IEF was perfomed on an 17-cm length of IPG strip having a pH range of 3-10 (linear pH gradient). The second separation was done using a 12% SDS-PAGE gel; A. control, B. cheongkukjang, C. control, D. soy E. genistein, F. control
 표
Ligand binding on FITC-labed protein of IEC cells. 콩이나 아몬드 단백질들은 BSA 와 비교할 때 현저하게 높은 결합을 보였으며 digested 또는 undigested 단백질들이 모두 결합됨을 알 수 있다 (A). 콩과 아몬드 단백질이 소장점막 단백질 중 32 kDa 단백질에 특이적으로 결합하였다 (B). Soy protein이 결합된 affinity column을 만들어서 장점막단백질과의 결합을 이루고 콩단백질과 결합하는 장점막단백질을 분리하여 1-D SDS-PAGE로 단백질을 분리하였다(C)
표
Ligand binding on FITC-labed protein of IEC cells. 콩이나 아몬드 단백질들은 BSA 와 비교할 때 현저하게 높은 결합을 보였으며 digested 또는 undigested 단백질들이 모두 결합됨을 알 수 있다 (A). 콩과 아몬드 단백질이 소장점막 단백질 중 32 kDa 단백질에 특이적으로 결합하였다 (B). Soy protein이 결합된 affinity column을 만들어서 장점막단백질과의 결합을 이루고 콩단백질과 결합하는 장점막단백질을 분리하여 1-D SDS-PAGE로 단백질을 분리하였다(C)
| 과제명(ProjectTitle) : | - |
|---|---|
| 연구책임자(Manager) : | - |
| 과제기간(DetailSeriesProject) : | - |
| 총연구비 (DetailSeriesProject) : | - |
| 키워드(keyword) : | - |
| 과제수행기간(LeadAgency) : | - |
| 연구목표(Goal) : | - |
| 연구내용(Abstract) : | - |
| 기대효과(Effect) : | - |
Copyright KISTI. All Rights Reserved.

※ AI-Helper는 부적절한 답변을 할 수 있습니다.