최소 단어 이상 선택하여야 합니다.
최대 10 단어까지만 선택 가능합니다.
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
NTIS 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
DataON 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Edison 바로가기
다음과 같은 기능을 한번의 로그인으로 사용 할 수 있습니다.
Kafe 바로가기
| 주관연구기관 | 한국과학기술연구원 Korea Institute Of Science and Technology |
|---|---|
| 연구책임자 | 양은경 |
| 참여연구자 | 공두식 , 이상원 , 김영수 , 백은옥 , 김현석 , 임영석 |
| 보고서유형 | 최종보고서 |
| 발행국가 | 대한민국 |
| 언어 | 한국어 |
| 발행년월 | 2017-05 |
| 과제시작연도 | 2016 |
| 주관부처 | 미래창조과학부 Ministry of Science, ICT and Future Planning |
| 과제관리전문기관 | 한국연구재단 National Research Foundation of Korea |
| 등록번호 | TRKO202000007174 |
| 과제고유번호 | 1711038088 |
| 사업명 | 포스트게놈신산업육성을위한다부처유전체(미래부) |
| DB 구축일자 | 2020-09-26 |
| 키워드 | 유전단백체.치료 불응 난치암.유전단백체-약물반응 프로파일.동반진단.HTS 항암치료반응 예측.Genome- proteome(Proteogenome).Refractory cancers.Proteogenomic-drug response profile.Companion diagnostics.HTS drug response prediction. |
- 악성 뇌종양 50례 및 진행성 폐암 22례에 대한 유전체, 전사체 및 종양유래세포를 이용한 60종 표적항암제의 약물 반응성을 분석하고, 해당 샘플에 대해 유전단백체-약물반응 랜드스케잎 구축을 지원함. 유전체-약물반응지도 기반의 바이오마커를 발굴하고 검증하고, 정밀의료를 위한 항암치료반응 예측 알고리즘을 개발하여 임상적용 플랫폼을 구축함
- 총 99명 환자의 내시경 조직에 대해 전사체, 글로벌단백체, 인산화단백체를 생산하고, 생산된 다중 다층 오믹스 데이터의 유전단백체 통합분석법을 구축/적용함
- 난치암 단백체의 오믹
- 악성 뇌종양 50례 및 진행성 폐암 22례에 대한 유전체, 전사체 및 종양유래세포를 이용한 60종 표적항암제의 약물 반응성을 분석하고, 해당 샘플에 대해 유전단백체-약물반응 랜드스케잎 구축을 지원함. 유전체-약물반응지도 기반의 바이오마커를 발굴하고 검증하고, 정밀의료를 위한 항암치료반응 예측 알고리즘을 개발하여 임상적용 플랫폼을 구축함
- 총 99명 환자의 내시경 조직에 대해 전사체, 글로벌단백체, 인산화단백체를 생산하고, 생산된 다중 다층 오믹스 데이터의 유전단백체 통합분석법을 구축/적용함
- 난치암 단백체의 오믹스 분석 및 생분자 시그너처 개발을 수행하고, 아바타 모델을 이용한 약물 반응 단백체를 발굴하고, 다중 표지 정량(MRM)을 이용한 난치암 임상적 예후를 검증하였으며, 유전단백체 신기술 개발 및 단백체 기반 난치암 분류 및 표적치료제 반응성 모델을 구축하고 유전단백체 통계 분석법 개발 및 데이터마이닝 소프트웨어를 구축함
- 난치암 감시진단을 위한 다중 마커 패널 확립 및 MRM Assay Platform을 구축하고, 간암 고위험군 감시진단을 위한 혈액 다중 마커 발굴 및 검증을 수행함. 또한, 위암 재발 모니터링 및 항암제 반응성 다중 마커를 개발함
(출처 : 보고서 요약서 3p)
ㅇ Purpose & Contents
Purpose:
In order to develop a drug library extension and drug reactivity prediction method for intractable cancer patients, target proteins were discovered by performing proteogenomic analysis from stomach cancer, lung cancer, and glioblastoma tissues. Those target protei
ㅇ Purpose & Contents
Purpose:
In order to develop a drug library extension and drug reactivity prediction method for intractable cancer patients, target proteins were discovered by performing proteogenomic analysis from stomach cancer, lung cancer, and glioblastoma tissues. Those target proteins were validated in blood samples with gastric cancer and liver cancer by MRM methods. We also applied the top-down analysis platform and the N-terminome discovery technology developed in the previous stage to the clinical samples.
Contents:
1. Establishment of The Cancer Proteogenomic Landscape and multiomics-guided drug response prediction algorithms through integrative analysis of proteogenome and HTS-based drug sensitivities of patient-derived cell models.
2. Application of proteogenomic integrative analysis platform in efforts to develop companion diagnosis methods for response prediction of chemotheraphy ro gastric cancer patients & use of the same platform to generate proteome data from the cancer samples from which PDX mouses were constructed.
3. Clinical MRM-MS analysis for more than 300 cases of malignant gastric cancer & discovery of lung cancer prediction markers through the proteogenomic method and verification of more than 50 cases of lung cancer sera.
4. Optimization of the multiplex MRM assay based on the surveillance marker candidates of gastric cancer remission (105 proteins, 158 peptides) and high-risk hepatocellular carcinoma (177 proteins, 388 peptides). The surveillance marker candidates of gastric cancer remission and high-risk hepatocellular carcinoma were tested via the multiplex MRM assay, yielding a result of AUC ≥ 0.8.
5. Improvement of top-down analysis platform and N-terminome discovery technologies established in Phase 1 for clinical sample application.
ㅇ Results
- Establishment of gene-drug association profiles by integrative analysis of cancer genome and drug responses against 60 targeted agents using patient-derived cell model (72 cases of malignant brain tumors and lung cancers).
- Gene-drug association profile-guided biomarker dicovery and validation (1 case)
- Establishment of gene-drug association map-guided precision cancer treatment platform
- For each of biopsy tumor tissue samples of 99 gastric cancer patients, a set of mRNA-seq, global quantitative proteome profile, and quantitative phosphoproteome profiling data was acquired using proteogenomic analysis platform.
- A standard operation protocol to obtain protein samples from small biopsy tumor tissues (less than 3 mg of wet tissue weight) was developed.
- From each of biopsy tumor tissue samples, more than 100,000 peptides and more than 10,000 phosphopeptides were identified with quantitative information.
- An advanced targeted MS platform was developed for high-throughput targeting on tissue samples.
- Analysis of global and phospho proteomes of brain tumor tissues (50 cases) to gain 9000 protein quantitative information and 9200 phosphorylation sites
- Clinical MRM-MS-mediated disease prognosis and prediction model were established through 227 cases of gastric cancer and 58 cases of lung cancer sera samples with biomarker candidates using plasma proteome analysis platform.
ㅇ Expected Contribution
- Extension of therapeutic option and suggestion of novel proteogenomic biomarker for precision diagnostics/treatment of refractory cancer patients
- Development of a proteogenomic prognosis method for gastric cancer
- Contribution to national competitiveness in new medical diagnosis market for gastric cancer
- Increase predication accuracy of drug response to refractory cancers by providing proteogenomics data to avatar mouse system
- Clinical applicability of proteomic-based refractory cancer subtype classification and target therapeutic agent accompanying diagnostic markers
- Development of a diagnostic method based on protein genome that minimizes the side effects of anticancer drugs and maximizes the effect
- Verified the validity of each multi-protein marker panel through large-scale clinical validation and constructed MRM assay platform to develop related industry and hospital linked service
- By performing data mining based on proteogenomics, we develop algorithms necessary for bioinformatics and contribute to the development of software architecture technology
- Multiple Marker Panel will be tested for efficacy in large scale clinical trials and developed into a Multiplex MRM assay platform that will be implemented in hospitals and related industries.
(출처 : SUMMARY 6p)
 표
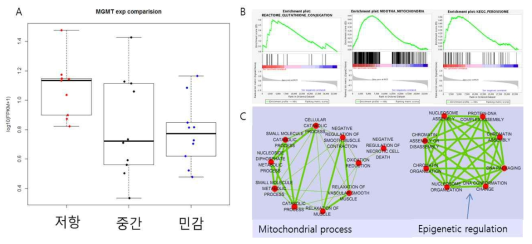
다중약물 저항 그룹에서 증가한 DEG 및 Gene set분석 결과. A 패널은 각 그룹별로 MGMT의 mRNA 발현량을 비교한 boxplot임 저항성 그룹이 민감 그룹에 비해 통계적으로 유의미하게 2배 가까이 증가해 있음. B 패널은 GSEA결과로 약물 동력학(pharmacokinetics)와 관련된 Gluthatione, Peroxsiome 외에 Mitochonria의 Gene set이 상대적으로 강화된 것으로 확인됨. C 패널은 GO-term 간의 연관성을 분석한 것으로 미토콘드리아 대사와 관련된 용어와 후성유전체적인 조절관련 용어로 다중약물 저항 그룹의 DEG들의 특징이 나타남을 보여줌
표
다중약물 저항 그룹에서 증가한 DEG 및 Gene set분석 결과. A 패널은 각 그룹별로 MGMT의 mRNA 발현량을 비교한 boxplot임 저항성 그룹이 민감 그룹에 비해 통계적으로 유의미하게 2배 가까이 증가해 있음. B 패널은 GSEA결과로 약물 동력학(pharmacokinetics)와 관련된 Gluthatione, Peroxsiome 외에 Mitochonria의 Gene set이 상대적으로 강화된 것으로 확인됨. C 패널은 GO-term 간의 연관성을 분석한 것으로 미토콘드리아 대사와 관련된 용어와 후성유전체적인 조절관련 용어로 다중약물 저항 그룹의 DEG들의 특징이 나타남을 보여줌
 표
다중약물 저항 그룹에서 증가한 DEG 및 Gene set분석 결과. A 패널은 각 그룹별로 MGMT의 mRNA 발현량을 비교한 boxplot임 저항성 그룹이 민감 그룹에 비해 통계적으로 유의미하게 2배 가까이 증가해 있음. B 패널은 GSEA결과로 약물 동력학(pharmacokinetics)와 관련된 Gluthatione, Peroxsiome 외에 Mitochonria의 Gene set이 상대적으로 강화된 것으로 확인됨. C 패널은 GO-term 간의 연관성을 분석한 것으로 미토콘드리아 대사와 관련된 용어와 후성유전체적인 조절관련 용어로 다중약물 저항 그룹의 DEG들의 특징이 나타남을 보여줌
표
다중약물 저항 그룹에서 증가한 DEG 및 Gene set분석 결과. A 패널은 각 그룹별로 MGMT의 mRNA 발현량을 비교한 boxplot임 저항성 그룹이 민감 그룹에 비해 통계적으로 유의미하게 2배 가까이 증가해 있음. B 패널은 GSEA결과로 약물 동력학(pharmacokinetics)와 관련된 Gluthatione, Peroxsiome 외에 Mitochonria의 Gene set이 상대적으로 강화된 것으로 확인됨. C 패널은 GO-term 간의 연관성을 분석한 것으로 미토콘드리아 대사와 관련된 용어와 후성유전체적인 조절관련 용어로 다중약물 저항 그룹의 DEG들의 특징이 나타남을 보여줌
| 과제명(ProjectTitle) : | - |
|---|---|
| 연구책임자(Manager) : | - |
| 과제기간(DetailSeriesProject) : | - |
| 총연구비 (DetailSeriesProject) : | - |
| 키워드(keyword) : | - |
| 과제수행기간(LeadAgency) : | - |
| 연구목표(Goal) : | - |
| 연구내용(Abstract) : | - |
| 기대효과(Effect) : | - |
Copyright KISTI. All Rights Reserved.

※ AI-Helper는 부적절한 답변을 할 수 있습니다.